Mastering CRISPRa and CRISPRi Screens: A Complete Guide to Gene Activation and Repression for Functional Genomics
This comprehensive guide provides researchers, scientists, and drug development professionals with essential knowledge and practical strategies for designing, executing, and interpreting CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screens.
Mastering CRISPRa and CRISPRi Screens: A Complete Guide to Gene Activation and Repression for Functional Genomics
Abstract
This comprehensive guide provides researchers, scientists, and drug development professionals with essential knowledge and practical strategies for designing, executing, and interpreting CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screens. Covering foundational principles, cutting-edge methodologies, common troubleshooting approaches, and validation techniques, this article synthesizes current best practices for leveraging these powerful tools to map gene function, identify therapeutic targets, and understand complex biological networks. The content addresses key intents from experimental design to data analysis, empowering researchers to implement robust screening workflows.
Understanding CRISPRa and CRISPRi: Core Principles, Components, and When to Choose Which Tool
Fundamental Mechanisms
CRISPRa and CRISPRi are derivative technologies of the CRISPR-Cas9 system, repurposed for precise transcriptional modulation without altering the underlying DNA sequence. Both systems utilize a catalytically "dead" Cas9 (dCas9) that retains its DNA-binding ability but lacks endonuclease activity. The core mechanistic distinction lies in the effector domains fused to dCas9.
CRISPR Interference (CRISPRi): For transcriptional repression, dCas9 is fused to a repressive effector domain. The most common is the Kruppel-associated box (KRAB) domain from human KOX1. When the dCas9-KRAB complex is guided to a target site, typically within ~200 bp downstream of the transcription start site (TSS), it induces heterochromatin formation via histone H3 lysine 9 trimethylation (H3K9me3), leading to stable gene silencing.
CRISPR Activation (CRISPRa): For transcriptional activation, dCas9 is fused to transcriptional activator domains. Simple activators like VP64 (a tetramer of the VP16 peptide) are weak. Advanced systems, such as the SunTag or synergistic activation mediator (SAM), recruit multiple copies of activators. The SAM system, for example, uses dCas9-VP64 alongside an engineered sgRNA scaffold that binds MS2-p65-HSF1 activator proteins, leading to robust recruitment of the cellular transcriptional machinery.
Application Notes
These tools are foundational for functional genomics screens to identify genes involved in specific phenotypes.
- CRISPRa Screens: Used to identify genes whose overexpression confers a selectable phenotype (e.g., drug resistance, cell proliferation, differentiation). They are invaluable for finding oncogenes, therapeutic targets, and genes that can reprogram cell states.
- CRISPRi Screens: Used to identify genes whose loss-of-function (knockdown) confers a phenotype. They offer a more uniform and complete repression than RNAi, with fewer off-target effects, and are ideal for identifying essential genes, tumor suppressors, and synthetic lethal interactions.
Table 1: Quantitative Comparison of CRISPRa and CRISPRi Systems
| Feature | CRISPRi (dCas9-KRAB) | CRISPRa (SAM System) |
|---|---|---|
| Primary Effector | KRAB repressive domain | VP64, p65, HSF1 activators |
| Typical Repression/Activation Fold-Change | 5- to 20-fold repression | 10- to 1,000-fold activation |
| Optimal Targeting Region | -50 to +300 bp relative to TSS | -200 to +1 bp upstream of TSS |
| Key Epigenetic Mark | Induces H3K9me3 (heterochromatin) | Induces H3K27ac (active chromatin) |
| Screen Applications | Loss-of-function, essentiality, suppressor | Gain-of-function, resistance, differentiation |
| Multiplexing Potential | High (multiple sgRNAs) | High, but larger effector size |
Protocols
Protocol A: Design and Cloning for a Pooled CRISPRa/i Screen
Objective: Clone a pooled lentiviral sgRNA library targeting your gene set of interest into the appropriate dCas9-effector backbone plasmid.
Materials: Plasmid backbone (e.g., lenti-sgRNA-MS2 for SAM, lenti-sgRNA for KRAB), pooled oligonucleotide library, BsmBI restriction enzyme, T4 DNA ligase, electrocompetent cells. Procedure:
- Design: Design 3-5 sgRNAs per gene, targeting the optimal window (see Table 1). Include non-targeting control sgRNAs.
- Digestion: Digest the backbone plasmid with BsmBI to remove the stuffer fragment.
- Annealing & Phosphorylation: Anneal the pooled oligos and phosphorylate using T4 PNK.
- Ligation: Ligate the annealed oligos into the digested backbone at a 10:1 insert:vector molar ratio.
- Transformation & Amplification: Electroporate the ligation product into E. coli, plate on large bioassay dishes, and harvest the pooled plasmids. Sequence to validate library representation.
Protocol B: Lentiviral Production & Cell Line Engineering
Objective: Generate lentivirus and create a stable cell line expressing the dCas9-effector.
Materials: HEK293T cells, packaging plasmids (psPAX2, pMD2.G), transfection reagent, target cells (e.g., K562, HeLa). Procedure:
- Stable dCas9 Cell Line: Co-transfect HEK293T with dCas9-VP64 (for SAM) or dCas9-KRAB plasmid and packaging plasmids. Harvest virus at 48h and 72h. Transduce target cells and select with blasticidin (10 µg/mL) for 7 days.
- Library Transduction: Produce lentivirus from the pooled sgRNA library plasmid. Transduce the stable dCas9-effector cell line at a low MOI (~0.3) to ensure most cells receive only one sgRNA. Select with puromycin (1-2 µg/mL) for 5-7 days. Maintain a coverage of >500 cells per sgRNA.
Protocol C: Screening and Next-Generation Sequencing (NGS) Analysis
Objective: Perform the phenotypic selection and identify enriched/depleted sgRNAs.
Materials: PCR purification kits, NGS platform (e.g., Illumina), selection agent (e.g., drug). Procedure:
- Phenotypic Selection: After puromycin selection (Day 0), split cells into experimental (e.g., +drug) and control (-drug) arms. Culture for 14-21 population doublings.
- Genomic DNA Extraction: Harvest ~1e7 cells from each arm at endpoint. Extract gDNA.
- sgRNA Amplification: Perform a two-step PCR to add Illumina adapters and barcodes to the integrated sgRNA sequence.
- NGS & Analysis: Sequence the PCR products. Align reads to the reference sgRNA library. Use algorithms like MAGeCK or BAGEL to calculate the log2 fold-change and statistical significance (p-value) for each sgRNA and gene.
Diagrams
Title: Core Mechanisms of CRISPRi and CRISPRa
Title: Pooled CRISPRa/i Screening Workflow
The Scientist's Toolkit
Table 2: Essential Research Reagents for CRISPRa/i Screens
| Reagent | Function | Example/Note |
|---|---|---|
| dCas9-Effector Plasmids | Provides the backbone for KRAB (i) or VP64/activator (a). | lenti-dCas9-KRAB, lenti-SAM (dCas9-VP64-MPH). |
| sgRNA Library Plasmid | Lentiviral vector for sgRNA expression. Contains puromycin resistance. | lenti-sgRNA-MS2 (for SAM), lentiGuide-Puro (for KRAB). |
| Lentiviral Packaging Plasmids | Required for producing virus particles. | psPAX2 (gag/pol), pMD2.G (VSV-G envelope). |
| Cell Line for Viral Production | High-transfectability line for making virus. | HEK293T, Lenti-X 293T. |
| Target Cell Line | The cells for the genetic screen. Must be transducible. | K562, HeLa, iPSCs, primary-like models. |
| Selection Antibiotics | For selecting stable integrants of dCas9 and sgRNA. | Blasticidin (dCas9), Puromycin (sgRNA). |
| NGS Library Prep Kit | To amplify and barcode sgRNA sequences from genomic DNA. | Illumina-compatible PCR kits. |
| Analysis Software | For quantifying sgRNA enrichment/depletion from NGS data. | MAGeCK, BAGEL, CRISPRcloud. |
CRISPR activation (CRISPRa) and interference (CRISPRi) screens have revolutionized functional genomics within gene activation and repression research. These screens rely on a catalytically dead Cas9 (dCas9) as a programmable DNA-binding scaffold. The targeted transcriptional outcome is determined by effector domains—activators or repressors—fused to dCas9. This article details the core system components, providing application notes and protocols essential for designing and executing robust CRISPRa/i screens, a critical methodology in modern drug discovery and target validation.
Core Component Specifications & Quantitative Data
Table 1: Common dCas9 Variants for CRISPRa/i
| dCas9 Variant | Origin | Key Mutations (Catalytic Inactivation) | Common Usage | Notes |
|---|---|---|---|---|
| dCas9 (S. pyogenes) | Streptococcus pyogenes | D10A, H840A | Standard CRISPRi, base for fusions | High DNA binding affinity; large size (~160 kDa). |
| dCas9 (S. aureus) | Staphylococcus aureus | N580A, D10A | Delivery via AAV, in vivo applications | Smaller size (~125 kDa); different PAM (NNGRRT). |
| dCas9-KRAB | S. pyogenes | D10A, H840A + KRAB fusion | Standard CRISPRi repression | KRAB domain directly fused for potent repression. |
Table 2: Activator Domains
| Domain | Origin | Typical Architecture | Approximate Fold Activation* | Notes |
|---|---|---|---|---|
| VP64 | Herpes Simplex Virus | 4x VP16 repeats | 2-10x | Mild activator; often used in synergistic combinations. |
| p65 | Human NF-κB | Transactivation domain | 3-15x | Synergizes with VP64; part of VPR and SAM systems. |
| Rta | Epstein-Barr Virus | Strong viral transactivator | 5-20x | Potent alone; part of VPR and SAM systems. |
| VPR Triad | Composite | VP64 + p65 + Rta fused to dCas9 | 20-300x | Highly potent activation across many cell types. |
| SAM (SunTag) | Scaffolded | dCas9-VP64 + MS2-p65-HSF1 | 100-1000x | Recruits multiple activators via scaffold; very high activation. |
Table 3: Repressor Domains
| Domain | Origin | Mechanism | Approximate Repression Efficiency* | Notes |
|---|---|---|---|---|
| KRAB | Human Kox1 | Recruits heterochromatin-forming complexes (e.g., SETDB1, HP1) | 5-100x (up to 90% knockdown) | Gold standard; represses within ~200 bp of TSS. |
| SID4x | Engineered | 4x fusion of the mSin3 interaction domain (SID) | 10-200x (up to 95% knockdown) | Potent synthetic repressor; recruits mSin3/HDAC complex. |
| Mxi1 | Human | Mad Max-interacting repressor domain | 3-50x | Alternative, less common than KRAB. |
*Fold change varies significantly based on target gene, genomic context, and cell type.
Detailed Experimental Protocols
Protocol 1: Design and Cloning of a dCas9-Effector Construct for CRISPRi
Objective: Clone the KRAB repressor domain into a lentiviral dCas9 expression vector. Materials: dCas9 backbone plasmid (e.g., pLV hU6-sgRNA hUbC-dCas9), KRAB domain PCR product, restriction enzymes (e.g., AgeI, EcoRI), T4 DNA Ligase, competent E. coli.
- Digestion: Set up digestion of the backbone plasmid and the KRAB insert with AgeI and EcoRI. Incubate at 37°C for 1 hour.
- Purification: Gel-purify the digested backbone and insert fragments.
- Ligation: Assemble a 20 µL ligation reaction with a 3:1 molar ratio of insert to backbone. Use T4 DNA Ligase and incubate at 16°C for 16 hours.
- Transformation: Transform 5 µL of the ligation mix into 50 µL of high-efficiency competent cells. Plate on selective antibiotic plates.
- Validation: Pick colonies, miniprep DNA, and verify the sequence by Sanger sequencing across the fusion junction.
Protocol 2: Lentiviral Production for CRISPRa Screen (SAM System)
Objective: Produce high-titer lentivirus for the dCas9-VP64 activator, MS2-p65-HSF1 activator, and sgRNA library. Materials: Lenti-X 293T cells, PEI transfection reagent, packaging plasmids (psPAX2, pMD2.G), SAM system plasmids (lenti dCas9-VP64, lenti MS2-p65-HSF1, sgRNA library plasmid), 0.45 µm PVDF filter.
- Day 0: Seed 10 million Lenti-X 293T cells in a 15 cm dish in DMEM + 10% FBS, no antibiotics.
- Day 1 (Transfection): For each virus (dCas9, activator, library), prepare DNA mix: 18 µg transfer plasmid, 12 µg psPAX2, 6 µg pMD2.G in 1.5 mL Opti-MEM. In a separate tube, mix 108 µL PEI with 1.5 mL Opti-MEM. Combine, vortex, incubate 15 min at RT, then add dropwise to cells.
- Day 2: Replace medium with 20 mL fresh, pre-warmed complete medium.
- Day 3 & 4: Harvest viral supernatant at 48 and 72 hours post-transfection. Pool harvests, filter through a 0.45 µm PVDF filter. Aliquot and store at -80°C or concentrate via ultracentrifugation.
Protocol 3: Performing a CRISPRi Knockdown Screen
Objective: Execute a pooled loss-of-function screen using dCas9-KRAB. Materials: Target cell line, lentivirus for dCas9-KRAB and sgRNA library, polybrene (8 µg/mL), puromycin, genomic DNA extraction kit, PCR primers for NGS library prep.
- Stable Cell Line Generation: Transduce target cells with dCas9-KRAB virus. Select with appropriate antibiotic (e.g., blasticidin) for 7-10 days.
- Library Transduction: Transduce dCas9-expressing cells with the sgRNA library virus at a low MOI (~0.3) to ensure single integration. Include 8 µg/mL polybrene. Spinoculate if needed.
- Selection and Passaging: 48 hours post-transduction, select with puromycin for 5-7 days to eliminate untransduced cells.
- Phenotype Application: Passage cells for the duration of the experiment (e.g., 14-21 days for a proliferation screen), maintaining library coverage of >500 cells per sgRNA.
- Genomic DNA (gDNA) Harvest: Harvest at least 50 million cells per replicate/timepoint. Extract gDNA.
- sgRNA Amplification & Sequencing: Perform a two-step PCR to amplify sgRNA cassettes from gDNA and add Illumina adaptors/indexes. Purify and pool libraries for next-generation sequencing.
- Analysis: Map sequencing reads to the sgRNA library. Use specialized algorithms (e.g., MAGeCK) to identify enriched or depleted sgRNAs under the selection condition.
Diagrams
Title: CRISPRi Gene Repression Mechanism via dCas9-KRAB
Title: CRISPRa Screen Workflow Using the SAM System
The Scientist's Toolkit
Table 4: Essential Research Reagent Solutions
| Item | Function/Application | Example Product/Catalog Number |
|---|---|---|
| Lentiviral dCas9-Effector Plasmids | Stable expression of dCas9 fused to activator/repressor domains. | Addgene: #61425 (dCas9-KRAB), #61426 (dCas9-VP64), #1000000078 (dCas9-VPR). |
| sgRNA Library Cloning Vector | Backbone for synthesizing and cloning pooled sgRNA libraries. | Addgene: #104875 (lentiGuide-Puro, for CRISPRi/a with MS2). |
| Second Activator Plasmid (for SAM) | Expresses the MS2-fused activator protein (p65-HSF1). | Addgene: #61427 (MS2-p65-HSF1). |
| Lentiviral Packaging Plasmids | Necessary for producing replication-incompetent lentiviral particles. | Addgene: #12260 (psPAX2), #12259 (pMD2.G). |
| Polybrene (Hexadimethrine Bromide) | Enhances viral transduction efficiency by neutralizing charge repulsion. | Sigma-Aldrich, H9268. |
| Puromycin Dihydrochloride | Selection antibiotic for cells transduced with puromycin-resistant vectors. | Thermo Fisher, A1113803. |
| Next-Generation Sequencing Kit | For preparing amplified sgRNA libraries for Illumina sequencing. | Illumina Nextera XT DNA Library Prep Kit. |
| Genomic DNA Extraction Kit | High-yield gDNA extraction from millions of cultured cells. | Qiagen Blood & Cell Culture DNA Maxi Kit. |
| Analysis Software | Statistical analysis of screen hits from NGS read counts. | MAGeCK (https://sourceforge.net/p/mageck/wiki/Home/). |
Traditional CRISPR-Cas9 knockout (KO) screens are powerful for identifying loss-of-function phenotypes but have significant limitations: they cannot study essential genes (as their complete knockout is lethal), cannot induce gain-of-function (GOF) phenotypes, and poorly control for hypomorphic (partial loss-of-function) effects. CRISPR activation (CRISPRa) and interference (CRISPRi) screens overcome these limitations by enabling tunable transcriptional modulation. This Application Note details protocols and frameworks for employing CRISPRa/i within functional genomic screens to explore these previously inaccessible biological spaces.
The table below summarizes the core capabilities of each screening modality.
Table 1: Comparison of CRISPR Screening Modalities
| Screening Modality | Primary Mechanism | Study Essential Genes? | Induce GOF? | Generate Hypomorphs? | Key Application |
|---|---|---|---|---|---|
| Traditional CRISPR-KO | Nuclease-induced DSBs, frameshift mutations | No (lethal) | No | Rare, stochastic | Complete loss-of-function |
| CRISPR Interference (CRISPRi) | dCas9 fused to repressive domains (e.g., KRAB) blocks transcription | Yes (titratable repression) | No | Yes (tunable) | Titratable knockdown, essential gene phenotyping |
| CRISPR Activation (CRISPRa) | dCas9 fused to activator domains (e.g., VPR, SAM) recruits transcriptional machinery | Yes (via overexpression) | Yes | No | Gene overexpression, suppressor screens |
Recent screen data (2023-2024) highlight the impact. For example, a genome-wide CRISPRi screen targeting essential genes in cancer cell lines achieved ~70-90% gene repression, identifying core fitness genes with a dynamic range of phenotypes impossible with KO. Parallel CRISPRa screens have identified resistance drivers with >10-fold gene activation.
Detailed Experimental Protocols
Protocol 1: CRISPRi Screen for Essential Gene Phenotyping & Hypomorphic Analysis
Objective: To identify and characterize dose-dependent phenotypes of essential genes using titratable repression. Reagents: Lentiviral CRISPRi library (e.g., Dolcetto or custom), HEK293T cells, polybrene (8 µg/mL), puromycin (2 µg/mL), doxycycline (for inducible systems). Workflow:
- Library Amplification & Virus Production: Amplify your chosen CRISPRi sgRNA library in E. coli with >=200x coverage. Co-transfect HEK293T cells with library plasmid, psPAX2, and pMD2.G using PEI. Harvest lentivirus at 48h and 72h post-transfection.
- Cell Line Engineering & Infection: Stably express dCas9-KRAB in your target cell line. Infect cells at a low MOI (<0.3) with library virus + polybrene. Select with puromycin for 7 days.
- Phenotypic Selection & Sampling: Passage cells, maintaining >=500x representation of each sgRNA. Collect genomic DNA from an initial time point (T0) and after phenotypic selection (e.g., 14-21 population doublings, or under drug challenge).
- Sequencing & Analysis: PCR-amplify integrated sgRNA sequences from gDNA. Sequence on an Illumina platform. Align reads to the library reference and quantify sgRNA abundance changes (e.g., using MAGeCK or PinAPL-Py). Hypomorphic phenotypes manifest as intermediate fold-changes compared to non-targeting controls.
Protocol 2: CRISPRa Screen for Gain-of-Function Phenotypes
Objective: To identify genes whose overexpression confers a selectable phenotype (e.g., drug resistance, proliferation). Reagents: Lentiviral CRISPRa library (e.g., Calabrese or SAM), target cell line, appropriate selection agent (e.g., chemotherapeutic). Workflow:
- Virus Production & Cell Infection: Follow steps similar to Protocol 1, using a CRISPRa library and a cell line expressing dCas9-VPR or the SAM complex components.
- Activation & Selection: After selection of transduced cells, apply the selective pressure (e.g., a low-dose drug). Maintain an unselected control population.
- Sample Processing: Harvest gDNA from selected and control populations after sufficient selection (e.g., when control population is ~50% confluent).
- Data Interpretation: Enriched sgRNAs in the selected population indicate genes whose activation drives resistance. Validation requires individual sgRNA or cDNA overexpression.
Visualizations
Diagram 1: Core Advantages of CRISPRa/i vs. KO Screens
Diagram 2: Experimental Decision Workflow for CRISPRa/i Screens
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions
| Item | Function & Specification |
|---|---|
| dCas9-KRAB Expression Vector | Constitutively or inductibly expresses nuclease-dead Cas9 fused to the KRAB transcriptional repressor domain. Core reagent for CRISPRi. |
| dCas9-VPR/SAM System | Expresses dCas9 fused to activator domains (VPR) or the synergistic activation mediator (SAM) complex components for robust CRISPRa. |
| Genome-wide sgRNA Libraries | Pre-designed pooled libraries (e.g., Dolcetto for CRISPRi, Calabrese for CRISPRa) targeting all human promoters with multiple sgRNAs/gene. |
| Lentiviral Packaging Plasmids | psPAX2 (gag/pol) and pMD2.G (VSV-G envelope) for production of replication-incompetent lentivirus. |
| Next-Generation Sequencing Kit | For high-throughput amplification and sequencing of sgRNA inserts from genomic DNA (e.g., Illumina Nextera XT). |
| Analysis Software (MAGeCK/PinAPL-Py) | Open-source tools for statistical analysis of screen data to identify significantly enriched or depleted sgRNAs/genes. |
| Polybrene | A cationic polymer that enhances viral transduction efficiency by neutralizing charge repulsion. |
| Puromycin/Selection Antibiotics | For selecting cells that have successfully integrated the sgRNA and effector construct. |
Primary Biological and Therapeutic Questions Addressed by Activation and Repression Screens
Within the broader thesis on CRISPRa (CRISPR activation) and CRISPRi (CRISPR interference) screening, these functional genomics tools address fundamental questions in biology and therapy. Activation screens (CRISPRa) systematically overexpress genes to identify those whose gain-of-function confers a phenotype, such as drug resistance or cell survival. Repression screens (CRISPRi) perform the opposite, knocking down gene expression to identify essential genes or sensitizers. Together, they map gene regulatory networks, identify therapeutic targets, and elucidate mechanisms of disease.
Key Biological & Therapeutic Questions
Table 1: Primary Questions Addressed by CRISPRa/i Screens
| Question Category | CRISPRa Application | CRISPRi Application | Example Therapeutic Goal |
|---|---|---|---|
| Gene Essentiality | Identify genes whose overexpression rescues cell death. | Identify genes whose loss causes cell death (essential genes). | Identify cancer cell vulnerabilities for targeted therapy. |
| Drug Mechanism & Resistance | Find genes causing drug resistance when overexpressed. | Find genes that sensitize to a drug when repressed (synthetic lethality). | Overcome chemotherapy resistance; identify combination therapies. |
| Disease Gene Discovery | Identify suppressors of disease phenotypes (e.g., toxin resistance). | Identify drivers of disease phenotypes (e.g., pathogen host factors). | Discover novel drug targets for genetic or infectious diseases. |
| Cellular Differentiation & Reprogramming | Identify transcription factors that drive lineage specification. | Identify genes that lock cells in a pluripotent state. | Develop protocols for regenerative medicine. |
| Signal Transduction Pathways | Identify pathway components that, when overexpressed, hyper-activate a pathway. | Identify negative regulators whose repression activates a pathway. | Target immune checkpoint pathways in oncology. |
Application Notes & Protocols
Application Note 1: Identifying Synthetic Lethal Interactions for Oncology
Objective: Use a CRISPRi screen to find genes whose repression synergistically kills cells with a specific oncogenic mutation. Biological Question: Which non-essential genes are synthetically lethal with mutant KRAS? Therapeutic Context: Developing targeted therapies for KRAS-mutant cancers.
Protocol: CRISPRi Synthetic Lethality Screen
- Cell Line Engineering: Generate a doxycycline-inducible dCas9-KRAB (for CRISPRi) HeLa cell line harboring mutant KRASG12C.
- Library Transduction: Transduce cells with a genome-wide CRISPRi lentiviral sgRNA library (e.g., human Brunello library) at low MOI (<0.3) to ensure single integration. Select with puromycin for 7 days.
- Screen Execution: Split cells into two arms: Control (Vehicle) and Treatment (KRAS inhibitor, e.g., Sotorasib). Maintain cells for 14-21 days, ensuring >200x coverage of each sgRNA.
- Genomic DNA Extraction & Sequencing: Harvest pellets, extract gDNA, PCR-amplify sgRNA regions, and sequence on an Illumina platform.
- Data Analysis: Use MAGeCK or BAGEL2 to compare sgRNA abundance between treatment and control. Identify sgRNAs depleted in the treatment arm, indicating genes whose repression sensitizes cells to KRAS inhibition.
Application Note 2: Discovering Novel Tumor Suppressors via CRISPRa
Objective: Use a CRISPRa screen to find genes whose overexpression inhibits tumor cell growth. Biological Question: Which genes, when activated, suppress proliferation in a glioblastoma model? Therapeutic Context: Gene therapy or targeted activation strategies.
Protocol: CRISPRa Positive Selection Growth Screen
- Cell Line Preparation: Use a glioblastoma stem cell (GSC) line stably expressing dCas9-VPR (CRISPRa activator).
- Library Transduction: Transduce with a CRISPRa sgRNA library targeting promoter regions of ~12,000 genes (e.g., SAM library). Select with blasticidin.
- Phenotypic Selection: Passage cells continuously for 4 weeks. Cells with growth-suppressive sgRNAs will be depleted over time.
- Sample Collection & NGS: Collect cells at Day 4 (T0) and Day 28 (Tfinal). Extract gDNA, prepare NGS libraries for sgRNA quantification.
- Data Analysis: Use MAGeCK or CRISPRcloud to identify sgRNAs significantly depleted in Tfinal vs. T0. The target genes of these sgRNAs are candidate tumor suppressors.
Visualization
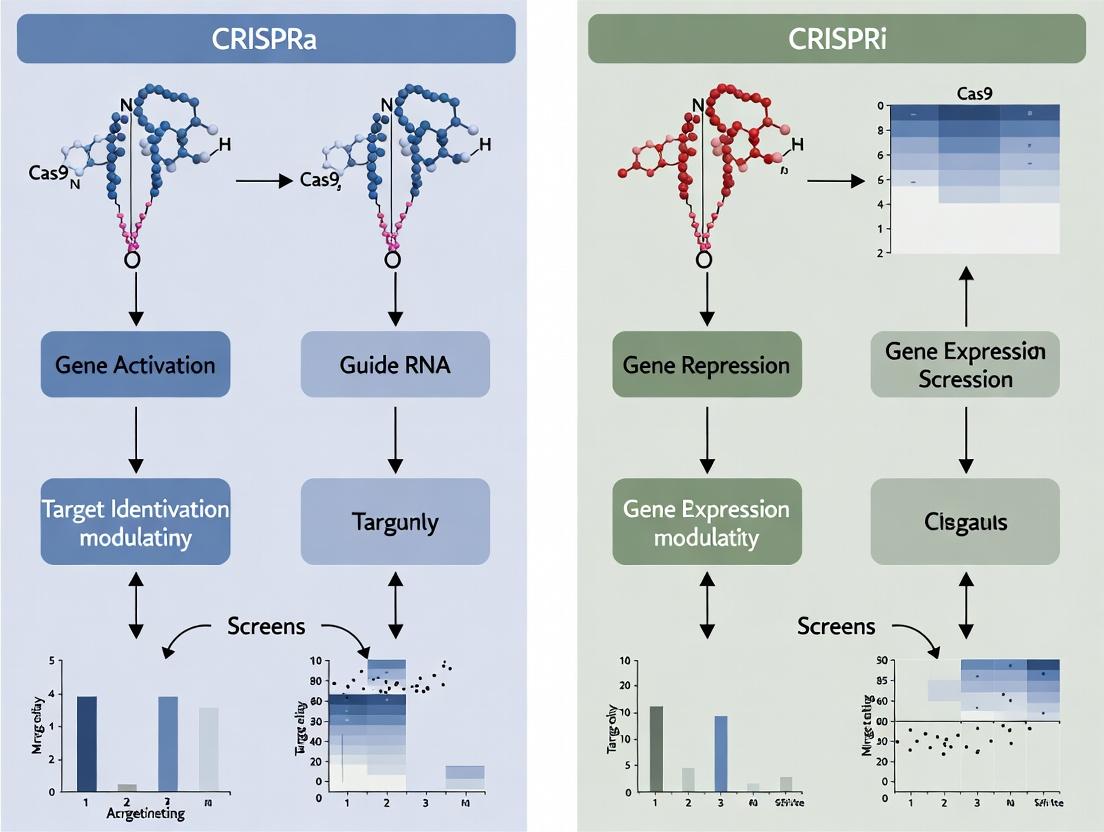
Diagram 1: CRISPRa vs. CRISPRi Core Mechanism
(Title: Core Mechanisms of CRISPRa and CRISPRi)
Diagram 2: Typical Workflow for a CRISPRa/i Screen
(Title: End-to-End Workflow for CRISPRa/i Genetic Screens)
Diagram 3: Key Questions in Drug Resistance Screens
(Title: Mapping Drug Resistance with CRISPRa/i)
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for CRISPRa/i Screens
| Reagent / Material | Function & Description | Example Product/ID |
|---|---|---|
| dCas9-Effector Plasmid | Stable expression vector for dCas9 fused to activator (VPR) or repressor (KRAB) domains. Essential for establishing the screening cell line. | lenti dCas9-VPR, lenti dCas9-KRAB |
| Genome-wide sgRNA Library | Pooled lentiviral library targeting all human genes with multiple sgRNAs per gene. Defines the screen's scope. | Brunello (CRISPRi), SAM (CRISPRa) |
| Lentiviral Packaging Mix | Plasmids (psPAX2, pMD2.G) for producing lentiviral particles of the sgRNA library in HEK293T cells. | psPAX2, pMD2.G |
| Selection Antibiotics | For selecting cells successfully transduced with library (puromycin) or dCas9-effector (blasticidin, puromycin). | Puromycin, Blasticidin S |
| Next-Generation Sequencing Kit | For preparing sequencing libraries from amplified sgRNA inserts. Critical for readout. | Illumina Nextera XT |
| Bioinformatics Software | Computational tools for analyzing NGS data to identify enriched/depleted sgRNAs and significant hits. | MAGeCK, BAGEL2, CRISPRcloud |
| Validated sgRNA Controls | Positive (essential gene) and negative (non-targeting) control sgRNAs for assay quality control. | e.g., PLKO anti-GFP sgRNA |
Within the broader thesis on CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screens for gene regulation research, the foundational step of experimental design dictates success. This application note details critical parameters for selecting sgRNA libraries, appropriate cell models, and expression systems to ensure robust, interpretable screens for drug target discovery and functional genomics.
sgRNA Library Selection and Design
The choice of library is paramount. Beyond simple gene coverage, considerations for CRISPRa/i screens include targeting specific transcriptional start sites (TSS) and avoiding confounding effects.
Key Design Principles:
- TSS Proximity: CRISPRi sgRNAs are most effective within -50 to +300 bp relative to the TSS. CRISPRa sgRNAs (e.g., using SAM or VPR systems) are typically placed within -400 to -50 bp upstream of the TSS.
- Specificity: Avoid off-targets by using optimized algorithms (e.g., from Doench et al., 2016) and recent genome builds.
- Redundancy: Libraries typically contain 3-10 sgRNAs per gene to mitigate sgRNA-specific inefficiencies.
- Control Guides: Essential sets include non-targeting controls (NTCs) and targeting essential/positive control genes.
Table 1: Comparison of Widely-Used CRISPRa/i Libraries
| Library Name | Primary Use | sgRNAs/Gene | Target Region | Key Feature | Common Expression System |
|---|---|---|---|---|---|
| Brunello (CRISPRko) | Knockout | 4 | Coding exons | High-efficiency, genome-wide | Lentiviral (lentiCRISPRv2) |
| Dolcetto | CRISPRi | 10 | -50 to +300 bp from TSS | Optimized for dCas9-KRAB | Lentiviral (pLV hU6-sgRNA hUbC-dCas9-KRAB) |
| Calabrese | CRISPRa | 10 | -400 to -50 bp from TSS | Optimized for SAM system | Lentiviral (lentiSAMv2) |
| CRISPRi-v2 (Horlbeck et al.) | CRISPRi | 3-5 | -50 to +300 bp from TSS | Compact, high-performance | Lentiviral (pHR-dCas9-KRAB-T2A-Puro) |
| SAM (Synergistic Activation Mediator) | CRISPRa | 3-5 | -400 to -50 bp from TSS | Uses MS2-p65-HSF1 activation domain | Lentiviral (lentiSAMv2, lentiMPHv2) |
Protocol 1.1: sgRNA Library Lentivirus Production (Lenti-X 293T Cell Method)
- Day 0: Seed Lenti-X 293T cells in poly-L-lysine coated 10-cm dishes at 5x10^6 cells/dish in DMEM + 10% FBS.
- Day 1: Transfect using a 3:1 ratio of PEI MAX (1 mg/mL) to total DNA. Per dish, combine:
- 4.5 µg Library sgRNA plasmid (e.g., lentiSAMv2)
- 3.0 µg psPAX2 (packaging plasmid)
- 1.5 µg pMD2.G (VSV-G envelope plasmid) in Opti-MEM. Incubate 15 min, add to cells.
- Day 2: Replace medium with 8 mL fresh, pre-warmed complete DMEM.
- Day 3 & 4: Harvest supernatant at 48h and 72h post-transfection. Pool harvests, centrifuge at 500 x g for 10 min, and filter through a 0.45 µm PES filter.
- Concentration: Concentrate virus 100x using Lenti-X Concentrator (Takara Bio) per manufacturer's instructions. Aliquot and store at -80°C.
- Titering: Determine functional titer (TU/mL) via puromycin selection or qPCR (Lenti-X qRT-PCR Titration Kit) on transduced HEK293T cells.
Cell Type Suitability and Engineering Requirements
Not all cell lines are equally amenable to CRISPRa/i screens. Key factors include proliferation rate, transducibility, and basal gene expression.
Table 2: Cell Line Suitability Assessment Criteria
| Criterion | Optimal for Screen | Potential Issue | Mitigation Strategy |
|---|---|---|---|
| Doubling Time | < 30 hours | Slow proliferation (< 48 hours) | Extend screen timeline; use metabolic (e.g., CellTiter-Glo) over proliferation assays. |
| Transduction Efficiency | > 80% (low MOI) | Low efficiency (< 40%) | Optimize polybrene/spinoculation; use alternative envelopes (e.g., RD114). |
| Ploidy | Diploid or Near-Diploid | Highly aneuploid/polyploid | Use more sgRNAs/gene; interpret copy-number effects cautiously. |
| Endogenous dCas9 Expression | None | N/A | Must engineer to express dCas9-activator/repressor fusion. |
| Proliferation Dependence on Target Pathway | Low | High | Use inducible dCas9 systems; short-term assay endpoints. |
Protocol 2.1: Generation of Stable dCas9-Expressing Cell Lines
- Pre-engineering: Transduce target cells with a lentivirus expressing the dCas9 fusion (e.g., dCas9-VPR for activation, dCas9-KRAB for repression) and a selectable marker (e.g., Blasticidin resistance). Use low MOI to avoid multiple integrations.
- Selection: Apply appropriate antibiotic (e.g., 5-10 µg/mL Blasticidin) for 7-10 days.
- Clonal Selection: Single-cell sort or limit dilute into 96-well plates. Expand clones.
- Validation:
- Western Blot: Confirm dCas9 fusion protein expression.
- Functional Test: Transduce with a positive control sgRNA (targeting a known highly activatable or repressible gene, e.g., CD69 or MYC) and measure gene expression via RT-qPCR 5-7 days post-transduction versus NTC.
Expression System Requirements
Sustained, stable, and balanced expression of both the dCas9-effector and the sgRNA is critical.
Core Requirements:
- Durability: Lentiviral integration ensures stable expression during long-term screens (e.g., 14+ days for proliferation).
- Moderate Expression: Very strong constitutive promoters (e.g., CMV) can lead to dCas9 toxicity. Use moderate promoters like EF1α or SFFV.
- Polycistronic Design: Link dCas9 to the selection marker via a self-cleaving peptide (e.g., T2A, P2A) for coordinated expression.
- sgRNA Expression: Use a pol III promoter (U6, H1). For dual-guide libraries (e.g., for SAM), two distinct pol III promoters are required.
Table 3: Essential Components of CRISPRa/i Expression Systems
| Component | CRISPRi Example | CRISPRa (SAM) Example | Function |
|---|---|---|---|
| dCas9 Fusion | dCas9-KRAB (repressor) | dCas9-VP64 (activator) | DNA-binding & transcriptional modulation |
| Effector Recruitment | N/A | MS2-p65-HSF1 (MCP fusion) | Synergistic activation (co-delivered or integrated) |
| dCas9 Promoter | EF1α | EF1α | Drives consistent, moderate dCas9 fusion expression |
| Selection Marker | Puromycin N-acetyl-transferase (PuroR) | Blasticidin S deaminase (BSD) | Selection for stable integrants |
| sgRNA Scaffold | Standard (for KRAB) | MS2 aptamer-modified | Bridges dCas9 and effectors (for SAM) |
Diagram 1: CRISPRa SAM System Assembly and Function (760px)
Diagram 2: CRISPRa i Screen Workflow Overview (760px)
The Scientist's Toolkit: Key Research Reagent Solutions
| Item / Reagent | Vendor Examples (Updated) | Function in CRISPRa/i Screens |
|---|---|---|
| Lentiviral sgRNA Library | Addgene (e.g., Dolcetto, Calabrese), Custom (Twist Bioscience) | Delivers gene-specific guides at scale for pooled screens. |
| dCas9-Effector Plasmids | Addgene (lentiSAMv2, pHR-dCas9-KRAB), Takara Bio | Source of transcriptional activator or repressor. |
| Lentiviral Packaging Mix | psPAX2/pMD2.G (Addgene), Lenti-X Packaging Single Shots (Takara Bio) | Required for producing replication-incompetent lentivirus. |
| Transfection Reagent | PEI MAX (Polysciences), Lipofectamine 3000 (Thermo Fisher) | For transient transfection of packaging cells during virus production. |
| Lentivirus Concentration Reagent | Lenti-X Concentrator (Takara Bio), PEG-it (System Biosciences) | Increases viral titer for hard-to-transduce cells. |
| Cell Line Engineering Kits | Lenti-X CRISPRa/i Kits (Takara Bio), DharmaFECT Transfection Reagents (Horizon) | Streamlines creation of stable dCas9-expressing cell lines. |
| Next-Gen Sequencing Library Prep Kit | NEBNext Ultra II DNA Library Prep (NGS), Illumina Kits | Prepares amplified sgRNA sequences for sequencing and abundance quantification. |
| Screen Analysis Software | MAGeCK (Broad), PinAPL-Py (IMBA), CRISPRAnalyzeR | Statistical analysis of screen data to identify significant hits. |
Step-by-Step Protocol: Designing and Executing a CRISPRa/i Screen from Library to Sequencing
CRISPR activation (CRISPRa) and interference (CRISPRi) screening technologies have revolutionized functional genomics, enabling systematic interrogation of gene function through targeted transcriptional modulation. Within this domain, two fundamental experimental design philosophies exist: hypothesis-driven and discovery-based (unbiased) screening. The choice of approach dictates screen design, library composition, analytical methods, and biological interpretation. This application note details the protocols, applications, and considerations for both strategies within CRISPRa/i research for drug discovery and target identification.
Table 1: Core Characteristics of Screening Approaches
| Feature | Hypothesis-Driven Screening | Discovery-Based Screening |
|---|---|---|
| Primary Goal | Test a specific biological hypothesis or mechanism. | Uncover novel genes/pathways without prior assumptions. |
| Library Design | Focused; genes selected based on prior knowledge (e.g., kinases, specific pathway). | Genome-wide or near-genome-wide coverage. |
| CRISPRa/i Library | Custom sub-libraries (e.g., focused activation of TF genes). | Standard genome-wide libraries (e.g., Calabrese, SAM, CRISPRi v2). |
| Experimental Throughput | Lower; manageable scale enables higher replication. | Very High; requires significant sequencing depth and resources. |
| Data Analysis | Simpler; often direct comparison of guide abundances. | Complex; requires robust normalization and hit-calling algorithms (MAGeCK, BAGEL). |
| Key Advantage | Deep mechanistic insight into a predefined system; higher signal-to-noise. | Unbiased discovery of novel regulators and unexpected biology. |
| Main Challenge | Limited to known biology; potential for confirmation bias. | High cost; high false-discovery rate; requires extensive validation. |
| Typical Hit Rate | Higher hit rate among tested genes. | Low hit rate, but absolute number of hits can be large. |
| Best For | Validating pathway models, probing drug mechanism-of-action, synthetic lethality. | Novel target identification, pathway discovery, phenotypically driven questions with no clear candidate genes. |
Table 2: Quantitative Comparison of Recent Published CRISPRa/i Screens (2023-2024)
| Study (PMID) | Approach | Library Size (guides) | Cell Model | Phenotype Readout | Primary Hits Identified |
|---|---|---|---|---|---|
| 38065903 | Discovery-based CRISPRa | ~70,000 (genome-wide) | iPSC-derived neurons | Neurite outgrowth | 120 significant gene activators |
| 38123654 | Hypothesis-driven CRISPRi | 5,000 (focused on chromatin regulators) | T cell leukemia | Resistance to chemotherapeutic | 8 key synthetic lethal regulators |
| 37935121 | Discovery-based CRISPRi | ~200,000 (genome-wide) | Macrophages | Inflammatory cytokine production | 45 repressors of IL-1β secretion |
| 38319877 | Hypothesis-driven CRISPRa | 1,200 (TF-focused sub-library) | Pancreatic cancer cells | Synergy with KRAS inhibitor | 3 transcription factors enhancing sensitivity |
Detailed Protocols
Protocol 1: Hypothesis-Driven CRISPRi Screen for Synthetic Lethality
Objective: Identify genes whose repression synergistically enhances cell death with a targeted oncology drug.
Research Reagent Solutions:
- Focused CRISPRi Lentiviral Library: Custom pooled library targeting 500 genes of interest (e.g., all DNA repair genes) with 10 sgRNAs/gene and 100 non-targeting controls.
- CRISPRi Stable Cell Line: Cell line expressing dCas9-KRAB-MeCP2 (e.g., HEK293T-HI) under antibiotic selection.
- Therapeutic Agent: The small-molecule inhibitor being studied (e.g., PARP inhibitor Olaparib).
- Next-Generation Sequencing (NGS) Kit: For amplifying and sequencing the integrated sgRNA region.
- Cell Viability Reagent: Such as CellTiter-Glo for ATP-based luminescent quantification.
Workflow:
- Library Amplification & Lentivirus Production: Amplify plasmid library in E. coli, prepare high-titer lentivirus using standard packaging plasmids (psPAX2, pMD2.G).
- Cell Infection & Selection: Infect CRISPRi cell line at a low MOI (~0.3) to ensure single integration. Select with puromycin for 7 days. Maintain a representation of >500 cells per sgRNA throughout.
- Treatment Arms: Split cells into two pools: A) DMSO vehicle control, B) Target drug (e.g., 1µM Olaparib).
- Phenotype Propagation: Culture cells for 14-21 days, allowing ~10-12 population doublings, to enrich for differential fitness effects.
- Genomic DNA Harvest & sgRNA Amplification: Harvest pellets of at least 50 million cells per arm at endpoint. Extract gDNA. Perform a two-step PCR to add sequencing adapters and sample barcodes to the sgRNA cassette.
- Sequencing & Analysis: Sequence on an Illumina platform. Align reads to the library reference. Use a tool like MAGeCK or BAGEL2 to compare sgRNA abundances between drug and control arms, identifying depleted sgRNAs (essential genes under drug treatment).
Title: Hypothesis-Driven CRISPRi Screen Workflow
Protocol 2: Discovery-Based CRISPRa Screen for Novel Resistance Genes
Objective: Unbiased identification of genes whose activation confers resistance to a cytotoxic compound.
Research Reagent Solutions:
- Genome-wide CRISPRa Library: e.g., Calabrese et al. (Nature, 2017) library (~70,000 sgRNAs targeting transcriptional start sites).
- CRISPRa Stable Cell Line: Cell line expressing MS2-p65-HSF1-dCas9-VP64 (SAM system) or dCas9-VPR.
- Selection Agent: The cytotoxic compound for which resistance mechanisms are sought.
- Deep Sequencing Reagents: Sufficient for >500x coverage of the library.
- Magnetic Beads for gDNA Cleanup: e.g., SPRIselect beads for PCR product purification.
Workflow:
- Library Infection & Selection: Infect CRISPRa cells at MOI ~0.3. Select with puromycin. Maintain >1000x representation.
- Baseline Sample (T0): Harvest 50 million cells pre-selection for gDNA as a reference.
- Positive Selection: Treat the remaining pool with a lethal dose (IC90) of the cytotoxic compound. Maintain culture, replenishing drug, until resistant population emerges (2-3 weeks).
- Endpoint Sample (T1): Harvest resistant population.
- NGS Library Prep: Isolate gDNA from T0 and T1. Perform large-scale PCR amplification of sgRNA region in multiple parallel reactions to avoid bias. Pool, purify, and quantify amplicons.
- Bioinformatic Analysis: Sequence to depth of >50 million reads per sample. Align to library. Use MAGeCK-MLE or PinAPL-Py to compare T1 vs T0, identifying significantly enriched sgRNAs/genes as resistance drivers.
Title: Discovery-Based CRISPRa Resistance Screen
The Scientist's Toolkit
Table 3: Essential Research Reagents for CRISPRa/i Screening
| Item | Function & Description | Example Product/Catalog |
|---|---|---|
| dCas9 Effector Plasmid | Stable expression of nuclease-dead Cas9 fused to transcriptional modulators (KRAB for i; VP64/p65/HSF1 for a). | pHAGE-dCas9-KRAB (Addgene #50919); lenti-dCas9-VPR (Addgene #63798). |
| Pooled sgRNA Library | Defined pool of lentiviral sgRNA constructs targeting the genome or a subset. | Human CRISPRa Calabrese Library (Addgene #1000000099); Human CRISPRi v2 (Addgene #83969). |
| Lentiviral Packaging Plasmids | For production of VSV-G pseudotyped lentivirus. | psPAX2 (Addgene #12260), pMD2.G (Addgene #12259). |
| Polycation Transfection Reagent | For efficient plasmid co-transfection in packaging cell line (HEK293T). | Polyethylenimine (PEI), Lipofectamine 3000. |
| Puromycin/Selection Agent | Selects for cells successfully transduced with the lentiviral library. | Puromycin dihydrochloride. |
| Next-Gen Sequencing Primer Sets | PCR primers for amplifying the sgRNA region and adding Illumina adapters/indexes. | Custom sequences per library design. |
| gDNA Extraction Kit | High-yield, high-quality genomic DNA extraction from cell pellets (min. 50-100µg). | Qiagen Blood & Cell Culture DNA Maxi Kit. |
| PCR Purification Beads | For size selection and clean-up of amplified sgRNA NGS libraries. | SPRIselect beads. |
| Bioinformatics Pipeline | Software for quantifying sgRNA reads and identifying significant hits. | MAGeCK (https://sourceforge.net/p/mageck), BAGEL2 (https://github.com/hart-lab/bagel). |
Within CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screening for gene function research, the selection and cloning of single guide RNA (sgRNA) libraries are foundational steps. This protocol details the considerations for choosing between genome-wide and focused libraries, outlines optimized design rules, and provides a method for high-efficiency library cloning, framed within the context of a thesis exploring transcriptional modulation screens for drug target discovery.
Library Selection: Genome-Wide vs. Focused
The choice between library types is dictated by the research question, resources, and downstream analysis capabilities.
Genome-Wide Libraries aim to target every gene in the genome. They are ideal for unbiased discovery and genome-scale functional genomics.
- Coverage: Typically 3-10 sgRNAs per gene.
- Size: 50,000 - 200,000+ sgRNAs.
- Application in Thesis: Essential for initial, hypothesis-generating screens to identify novel genes involved in a pathway or phenotype of interest for activation or repression.
Focused/Subset Libraries target a predefined set of genes (e.g., a pathway, gene family, or set of hits from a prior screen).
- Coverage: Higher density, often 5-10 sgRNAs per gene.
- Size: 100 - 10,000 sgRNAs.
- Application in Thesis: Optimal for hypothesis-driven secondary validation, mechanistic studies, or screens focusing on specific drug target classes (e.g., all kinases or epigenetic modifiers).
Quantitative Comparison Table
| Feature | Genome-Wide Library | Focused Library |
|---|---|---|
| Primary Goal | Unbiased discovery | Targeted validation/hypothesis testing |
| Typical Size | 50k - 200k+ sgRNAs | 100 - 10k sgRNAs |
| sgRNAs per Gene | 3-10 | 5-10 (or more) |
| Screen Cost | High | Moderate to Low |
| Sequencing Depth | High (≥ 500x) | Lower (≥ 200x) |
| Data Complexity | High, requires robust hit-calling | Lower, more straightforward analysis |
| Best for Thesis Stage | Initial discovery chapter | Validation & mechanistic follow-up chapters |
Optimized sgRNA Design Rules for CRISPRa/i
Effective design is critical for minimizing off-target effects and maximizing on-target efficacy in transcriptional modulation.
Core Design Parameters:
- Target Region: For CRISPRi, sgRNAs should target the transcription start site (TSS) or early exon, typically within -50 to +300 bp relative to the TSS. For CRISPRa, target the upstream promoter region, -400 to -50 bp from the TSS.
- Specificity: Minimize off-targets by ensuring ≤3-4 mismatches in the seed region (PAM-proximal 12 bases) across the genome.
- GC Content: Optimize between 40-60%.
- Avoidance Regions: Exclude sgRNAs with homopolymer runs (>4 bases), self-complementarity (which can affect expression), and SNPs.
Quantitative Design Rules Table
| Parameter | Optimal Value/Range | Rationale |
|---|---|---|
| CRISPRi Target Window | -50 to +300 bp from TSS | Optimal for dCas9-KRAB-mediated repression |
| CRISPRa Target Window | -400 to -50 bp from TSS | Optimal for dCas9-VPR/VP64 recruitment |
| sgRNA Length | 20 nt (spacer) | Standard for SpCas9 compatibility |
| GC Content | 40% - 60% | Balances stability and specificity |
| Seed Region Mismatches | ≥3-4 mismatches required for any potential off-target | Ensures high specificity |
| Number of sgRNAs/Gene | 3-5 (minimum), 5-10 (recommended) | Accounts for variable efficacy; enables robust statistics |
Protocol: Cloning an sgRNA Library into a Lentiviral Vector
This protocol describes bulk cloning of an oligonucleotide library into a lentiviral sgRNA expression backbone (e.g., lentiGuide-Puro, plentiCRISPRv2 with modifications for CRISPRa/i).
Materials & Reagents
Research Reagent Solutions Toolkit
| Item | Function |
|---|---|
| Designed Oligo Library Pool (ssDNA) | Contains the variable sgRNA spacer sequences flanked by vector-specific cloning overhangs. |
| BsmBI-v2 Restriction Enzyme (NEB) | Type IIS enzyme used for golden gate assembly; cuts outside its recognition sequence to generate unique overhangs. |
| T4 DNA Ligase & Buffer | Catalyzes the ligation of digested vector and insert fragments. |
| High-Capacity Lentiviral Backbone (e.g., pLV-sgRNA) | Contains BsmBI sites, sgRNA scaffold, mammalian/H1 promoter, and bacterial resistance marker. |
| NEB Stable Competent E. coli | High-efficiency cells for transformation of large, complex libraries to maintain diversity. |
| QIAprep Spin Miniprep Kit | For small-scale plasmid purification from individual colonies for quality control. |
| Maxiprep or Megaprep Kit | For large-scale plasmid DNA purification of the entire library pool for lentivirus production. |
| Next-Generation Sequencing (NGS) Kit (e.g., Illumina MiSeq) | For quantifying library representation and integrity. |
Detailed Protocol
Part A: Preparation of Vector and Insert
- Digest Backbone: Digest 5 µg of lentiviral sgRNA vector with BsmBI (or other appropriate Type IIS enzyme) according to manufacturer instructions. Gel-purify the linearized backbone.
- Prepare Insert: The synthesized oligonucleotide library is delivered as single-stranded DNA. Perform a limited-cycle PCR to amplify the library and add full BsmBI sites. Purify the PCR product.
Part B: Golden Gate Assembly
- Set up the Golden Gate reaction in a 20 µL volume:
- 50 ng BsmBI-digested, gel-purified vector
- 20 ng PCR-amplified insert library (molar ratio ~1:3 vector:insert)
- 1 µL BsmBI-v2 enzyme
- 1 µL T4 DNA Ligase
- 2 µL 10x T4 DNA Ligase Buffer
- Nuclease-free water to 20 µL.
- Run the following thermocycler program:
- (37°C for 5 min → 16°C for 5 min) x 25 cycles
- 37°C for 15 min
- 80°C for 15 min
- Hold at 4°C.
Part C: Transformation and Library Amplification
- Desalt the entire assembly reaction using a spin column.
- Electroporate the entire desalted product into NEB Stable Competent E. coli. Use ten 50 µL aliquots of cells to ensure >1000x library coverage.
- Pool all transformations, add SOC medium, and recover with shaking for 1 hour at 37°C.
- Plate the entire culture on five large (245 mm x 245 mm) LB agar plates with appropriate antibiotic. Incubate at 32°C for 18-24 hours. Lower temperature helps prevent recombination.
Part D: Harvesting and Validation
- Scrape all colonies and perform a Maxiprep to harvest the plasmid library. Determine concentration.
- Quality Control by NGS:
- Amplify the sgRNA insert region from 100 ng of the final plasmid library using primers containing Illumina adapters and sample barcodes.
- Purify and run on a MiSeq (or similar) system to obtain at least 500 reads per sgRNA expected in the library.
- Analyze data to confirm >90% of designed sgRNAs are present with even representation (no sgRNA should be overrepresented by >100-fold compared to the median).
Visualizations
sgRNA Library Selection and Cloning Workflow
sgRNA Target Windows for CRISPRi vs CRISPRa
Viral Delivery (Lentivirus) and Stable Cell Line Generation for dCas9 Effector Expression
Within the broader thesis on CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screens for gene regulation research, the generation of stable cell lines expressing the catalytically dead Cas9 (dCas9) effector is a critical foundational step. Stable expression ensures uniform and sustained levels of dCas9 fused to transcriptional activation (e.g., VPR, p65AD) or repression (e.g., KRAB, SID4x) domains, which is paramount for consistent, genome-wide screening outcomes. Lentiviral delivery remains the gold standard for this purpose due to its ability to transduce both dividing and non-dividing cells with high efficiency and stable genomic integration.
This application note details a streamlined protocol for producing high-titer lentivirus encoding dCas9 effectors and subsequently generating polyclonal stable cell lines, optimized for downstream CRISPRa/i screening workflows.
Key Research Reagent Solutions
The following table summarizes essential reagents and their functions in this protocol.
| Reagent/Category | Example/Description | Primary Function |
|---|---|---|
| dCas9 Effector Plasmid | lenti-dCas9-VPR, lenti-dCas9-KRAB | Source of the dCas9-transcriptional regulator fusion gene. Must be in a lentiviral backbone (e.g., pLX_311). |
| Lentiviral Packaging Plasmids | psPAX2, pMD2.G (3rd Gen) | psPAX2 provides gag, pol, rev; pMD2.G provides VSV-G envelope protein for viral particle production. |
| Transfection Reagent | Polyethylenimine (PEI), Lipofectamine 3000 | Facilitates DNA plasmid delivery into packaging cells (HEK293T). |
| Packaging Cell Line | HEK293T/17 | High transfection efficiency, robust viral particle production, and SV40 T-antigen for plasmid replication. |
| Target Cell Line | K562, HEK293, iPSCs, etc. | Cell line of interest for the eventual CRISPR screen. Must be amenable to lentiviral transduction. |
| Selection Antibiotic | Puromycin, Blasticidin | Selects for cells that have stably integrated the viral construct, based on the resistance gene on the transfer plasmid. |
| Transduction Enhancer | Polybrene (Hexadimethrine bromide) | Neutralizes charge repulsion between viral particles and cell membrane, increasing transduction efficiency. |
| Titer Quantification Reagent | qPCR Lentiviral Titer Kit | Quantifies functional viral vector genomes per mL (vg/mL) for accurate MOI calculation. |
Table 1: Typical Viral Production and Transduction Metrics
| Parameter | Typical Range/Value | Notes & Impact on Experiment |
|---|---|---|
| Transfection Efficiency (HEK293T) | >80% | Critical for high-titer virus. Assessed via fluorescent reporter co-transfection. |
| Functional Viral Titer (qPCR) | 1x10^7 - 1x10^9 vg/mL | Aim for >1x10^8 vg/mL for robust stable line generation. Concentration may be required. |
| Multiplicity of Infection (MOI) Used | 1 - 5 | For stable integration, a low MOI (e.g., MOI=1-3) is preferred to minimize multiple integrations. |
| Transduction Efficiency (Target Cells) | 30-95% | Depends on cell type. Use a fluorescent reporter virus to assess pre-selection. |
| Selection Duration | 5-10 days | Until all un-transduced control cells are dead. Cell type-dependent. |
| dCas9 Expression Validation (Western Blot) | N/A | Confirm expected protein size (~160 kDa for dCas9 plus fusion domain). |
| Functional Validation (qPCR) | 10-1000x fold change | Test known target gene activation (CRISPRa) or repression (CRISPRi) with guide RNA controls. |
Table 2: Advantages & Limitations of Lentiviral Stable Line Generation
| Aspect | Advantages | Limitations & Mitigations |
|---|---|---|
| Expression Stability | Consistent, long-term dCas9 effector expression over passages. | Potential for epigenetic silencing; use of S/MAR elements or routine antibiotic maintenance. |
| Cell Type Applicability | Broad (dividing & non-dividing, primary cells, stem cells). | Some cell types (e.g., primary B cells) remain difficult; optimize enhancers/spinoculation. |
| Uniformity | Polyclonal population averages out position effects. | Clonal variation exists; use polyclonal pools with high representation (>200 independent clones). |
| Safety | 3rd generation, split-packaging, VSV-G pseudotyped vectors are biosafety level 2. | Requires appropriate biosafety containment for production and handling. |
| Time Investment | Once established, process is scalable and reproducible. | Initial virus production and selection requires 3-4 weeks. |
Detailed Protocols
Protocol 4.1: High-Titer Lentivirus Production (PEI Transfection in HEK293T)
Objective: Produce replication-incompetent lentivirus encoding dCas9-VPR or dCas9-KRAB.
Materials:
- HEK293T cells (low passage, >90% viability)
- High-glucose DMEM with GlutaMAX, 10% FBS, 1% Pen/Strep
- Opti-MEM Reduced Serum Medium
- Plasmids: Transfer plasmid (dCas9-effector), psPAX2, pMD2.G
- Linear PEI (1 mg/mL in water, pH 7.0)
- 0.45 μm PVDF syringe filter
- 10 cm tissue culture dishes or 10-layer cell factories for scale-up
Method:
- Day 0: Plate Cells. Seed HEK293T cells at ~3-4 x 10^6 cells per 10 cm dish in 10 mL complete DMEM. Incubate overnight at 37°C, 5% CO2. Target ~70-80% confluency at time of transfection.
- Day 1: Transfect.
- Prepare DNA Mix per dish: 10 μg transfer plasmid + 7.5 μg psPAX2 + 2.5 μg pMD2.G in 500 μL Opti-MEM.
- Prepare PEI Mix per dish: 45 μL PEI (1 mg/mL) in 500 μL Opti-MEM. Vortex briefly.
- Combine DNA and PEI mixes. Vortex immediately for 15 sec. Incubate at room temp for 15-20 min.
- Add the 1 mL DNA:PEI complex dropwise to the dish. Gently rock the dish.
- Return to incubator.
- Day 2: Refresh Media. ~16 hours post-transfection, carefully aspirate media and replace with 10 mL fresh, pre-warmed complete DMEM.
- Day 3 & 4: Harvest Virus. 48 and 72 hours post-transfection, collect the virus-containing supernatant. Pass through a 0.45 μm PVDF filter to remove cell debris. Pool harvests if desired.
- Optional: Concentrate virus using PEG-it or ultracentrifugation. Resuspend pellet in PBS or medium.
- Aliquot, snap-freeze in liquid nitrogen or dry ice/ethanol, and store at -80°C. Avoid repeated freeze-thaw cycles.
Protocol 4.2: Generation of Polyclonal Stable dCas9 Effector Cell Line
Objective: Transduce target cells and select a polyclonal population stably expressing the dCas9 effector.
Materials:
- Target cells (e.g., K562, HEK293)
- Appropriate growth medium
- Lentiviral supernatant (from Protocol 4.1)
- Polybrene (stock 4-8 mg/mL in water)
- Appropriate selection antibiotic (e.g., Puromycin, Blasticidin)
- Crystal violet or cell viability stain (for kill curve)
Method:
- Determine Selection Kill Curve: Prior to transduction, perform a kill curve on untransduced target cells with the antibiotic (e.g., 0.5-10 μg/mL puromycin). The minimal concentration that kills all cells in 5-7 days is the working concentration.
- Day 0: Transduce.
- Seed target cells at ~50% confluency (or 2-5x10^5 cells/mL for suspension cells) in a 6-well plate.
- Prepare transduction mix: Fresh medium + viral supernatant (volume based on desired MOI and titer) + Polybrene (final concentration 4-8 μg/mL). Include a no-virus control well.
- Replace cell media with the transduction mix. For suspension cells, spinoculate (centrifuge at 800-1000 x g for 30-90 min at 32°C) to enhance infection.
- Incubate for 24 hours.
- Day 1: Remove Virus. Aspirate transduction mix from adherent cells (or centrifuge and resuspend suspension cells) and replace with fresh, complete growth medium.
- Day 2: Begin Selection. Replace medium with fresh medium containing the predetermined concentration of selection antibiotic.
- Days 3-10: Maintain Selection. Refresh antibiotic-containing medium every 2-3 days. Monitor cell death in the control well. Continue selection until all control cells are dead and transduced wells show healthy, proliferating cells.
- Expand and Validate: Expand the polyclonal stable pool. Validate dCas9 effector expression by Western blot and functional assays (e.g., using a validated sgRNA and qPCR for target gene expression change).
- Bank Cells: Cryopreserve multiple vials of the validated stable pool for future screening use.
Visualizations
Title: Lentiviral Production Workflow
Title: dCas9-Effector Pathways in CRISPRa/i Screens
Title: Stable dCas9 Cell Line Generation Protocol
CRISPR activation (CRISPRa) and interference (CRISPRi) screens are powerful tools for identifying genes that modulate specific cellular phenotypes when their expression is systematically perturbed. The execution phase—encompassing the transduction of guide RNA (gRNA) libraries, selection of successfully modified cells, and application of a defined phenotypic pressure—is critical for screen success. This protocol details the methodology for performing pooled CRISPRa/i screens, focusing on the application of selective pressures such as drug treatment or survival assays to uncover genetic regulators of drug response, cellular fitness, and survival pathways. These screens are integral to target discovery and validation in therapeutic development.
Key Research Reagent Solutions
| Reagent/Material | Function in Screen Execution |
|---|---|
| Lentiviral gRNA Library | Delivers CRISPRa (e.g., SAM, VPR) or CRISPRi (dCas9-KRAB) machinery and sequence-specific gRNAs to cells for pooled genetic perturbation. |
| Polybrene (Hexadimethrine Bromide) | A cationic polymer that enhances viral transduction efficiency by neutralizing charge repulsion between virions and cell membranes. |
| Puromycin / Blasticidin | Selection antibiotics used to eliminate untransduced cells following lentiviral library delivery, ensuring a pure population of CRISPR-modified cells. |
| Phenotypic Pressure Agent (e.g., Drug Compound) | The applied selective condition (e.g., IC50-IC90 concentration of a therapeutic agent) to challenge the cell population and enrich for gRNAs conferring resistance or sensitivity. |
| Cell Viability Stain (e.g., Propidium Iodide) | Used in FACS-based survival screens to distinguish and isolate live vs. dead cell populations for downstream sequencing. |
| Genomic DNA Extraction Kit | For high-yield, high-quality gDNA isolation from screen samples prior to gRNA amplification and sequencing. |
| PCR Primers for gRNA Amplification | Flanking primers containing Illumina adapter sequences to amplify the integrated gRNA cassette from genomic DNA for NGS library preparation. |
| SPRI Beads | For size selection and purification of PCR-amplified gRNA libraries, removing primers and primer dimers. |
Detailed Experimental Protocol
Protocol Part A: Library Transduction and Selection
Objective: To generate a population of cells with comprehensive genomic perturbations at high coverage.
- Day -1: Cell Plating: Plate the target cells (e.g., HEK293T, K562, A549) in antibiotic-free growth medium. Seed enough cells to achieve 30-40% confluence on the day of transduction.
- Day 0: Viral Transduction:
- Calculate the required volume of lentiviral library supernatant to achieve a low Multiplicity of Infection (MOI ~0.3-0.4) and a final library coverage of >500 cells per gRNA. Include polybrene at a final concentration of 4-8 µg/mL.
- Replace cell medium with the virus-polybrene mixture.
- Centrifuge plates at 800 x g for 30-60 minutes at 32°C (spinoculation) to enhance infection.
- Incubate at 37°C, 5% CO₂ for 6-24 hours.
- Day 1: Media Change: Carefully remove viral supernatant and replace with fresh growth medium.
- Days 2-5: Antibiotic Selection:
- Begin selection with the appropriate antibiotic (e.g., 1-5 µg/mL puromycin) 48 hours post-transduction.
- Maintain selection for 3-5 days, or until all cells in a non-transduced control well are dead.
- Harvest a sample of selected cells (~5x10⁶ cells). This is the T0 sample (baseline reference). Pellet, wash with PBS, and store at -80°C for gDNA extraction.
Protocol Part B: Application of Phenotypic Pressure
Objective: To challenge the perturbed cell population to enrich for gRNAs that alter the phenotype of interest.
- Day 5+: Split and Apply Pressure:
- Split the selected cell population into two arms: Experimental Pressure and Control (no pressure). Maintain each arm at a minimum of 500x library coverage.
- For a drug screen: Treat the experimental arm with the compound at a predetermined inhibitory concentration (e.g., IC70-IC90). The control arm receives vehicle (e.g., DMSO).
- For a survival/proliferation screen: The "pressure" is simply continued passaging; gRNAs affecting fitness will be depleted or enriched over time.
- Phenotype Execution:
- Drug Treatment: Culture cells under drug/vehicle pressure for 7-21 days, passaging as needed and replenishing the drug/vehicle.
- FACS-Based Survival: After a cytotoxic insult, stain cells with a viability dye (e.g., propidium iodide). Use FACS to isolate the top/bottom 10-20% of live cells. Pellet and freeze sorted populations.
- Endpoint Sampling: Harvest a minimum of 5x10⁶ cells from each experimental and control arm at the endpoint. Pellet, wash with PBS, and store at -80°C. This is the Tend sample.
Protocol Part C: gRNA Recovery and Sequencing
- Genomic DNA Extraction: Isolate gDNA from all frozen cell pellets (T0, ControlTend, ExperimentalTend) using a large-scale gDNA extraction kit. Quantify DNA precisely.
- gRNA Amplification & NGS Library Prep:
- Perform a two-step PCR. PCR1 amplifies the gRNA region from gDNA using primers containing partial Illumina adapters.
- Purify PCR1 products using SPRI beads.
- PCR2 adds full Illumina adapters and sample-specific barcodes.
- Purify the final library, quantify, and pool samples for sequencing on an Illumina platform (MiSeq/NextSeq/Novaseq). Aim for >100 reads per gRNA per sample.
Table 1: Critical Screen Parameters and Typical Values
| Parameter | Recommended Value | Rationale |
|---|---|---|
| Library Coverage (Cells/gRNA) | >500 | Ensures statistical robustness and minimizes stochastic dropout. |
| Multiplicity of Infection (MOI) | 0.3 - 0.4 | Limits multiple gRNA integrations per cell, ensuring a single perturbation per cell. |
| Antibiotic Selection Duration | 3 - 5 days | Ensures complete death of non-transduced cells without over-stressing the pool. |
| Minimum Cell Number for gDNA | 5 x 10⁶ | Yields sufficient gDNA for robust PCR amplification of the gRNA library. |
| Sequencing Depth (Reads/gRNA) | >100 | Provides accurate measurement of gRNA abundance changes. |
Table 2: Example Enrichment/Depletion Metrics from a Hypothetical Drug Resistance Screen
| gRNA Target Gene | Log2 Fold Change (Drug/Control) | p-value (adjusted) | Interpretation |
|---|---|---|---|
| ABCG2 | +3.85 | 1.2e-10 | Strongly enriched; known multidrug resistance transporter. |
| TP53 | -2.91 | 5.7e-08 | Strongly depleted; loss may enhance drug sensitivity. |
| Non-Targeting Ctrl_1 | +0.12 | 0.78 | Neutral control, shows no significant change. |
| Non-Targeting Ctrl_2 | -0.08 | 0.82 | Neutral control, shows no significant change. |
Visualizations
Title: Workflow for Pooled CRISPRa/i Phenotypic Screen
Title: Logic of Phenotype Induction & Screen Readout
Harvesting and Sample Preparation for Next-Generation Sequencing (NGS)
Within the thesis research focusing on CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screens for systematic gene regulation studies, high-quality NGS library preparation is the critical endpoint. These screens generate complex pools of genetically perturbed cells. The harvested genomic material contains the integrated guide RNA (gRNA) sequences, which serve as barcodes reflecting the genetic perturbation and its phenotypic outcome. Precise harvesting and sample preparation are paramount to accurately deconvolute screening results, linking gRNA abundance to gene activation or repression phenotypes.
Key Research Reagent Solutions
Table 1: Essential Reagents and Kits for NGS Sample Prep from CRISPR Screens
| Reagent/Kits | Function in Workflow |
|---|---|
| Cell Lysis Buffer (Proteinase K) | Digests cellular proteins and nucleases, releasing genomic DNA (gDNA) containing the integrated gRNA cassette. |
| Magnetic Beads (SPRI) | Size-selects and purifies DNA fragments (e.g., post-PCR), enabling cleanup and buffer exchange. |
| High-Fidelity PCR Master Mix | Amplifies the gRNA target region from genomic DNA with minimal bias and errors for accurate representation. |
| Unique Dual-Index (UDI) PCR Primers | Attaches sample-specific barcodes and Illumina sequencing adapters during PCR, enabling multiplexing. |
| Qubit dsDNA HS Assay Kit | Precisely quantifies low-concentration DNA samples (e.g., final libraries) for accurate pooling. |
| TapeStation/ Bioanalyzer HS DNA Kit | Assesses library fragment size distribution and quality, ensuring correct insert size. |
Experimental Protocols
Protocol 3.1: Harvesting Genomic DNA from CRISPR Pooled Screen Cells
Objective: To isolate high-quality, high-molecular-weight gDNA from fixed or pelleted cell populations.
Materials: Cell pellet (> 1e6 cells), PBS, Proteinase K, Lysis Buffer, Isopropanol, Ethanol (70%), TE Buffer.
Procedure:
- Cell Lysis: Resuspend cell pellet in 500 µL of PBS. Add 500 µL of Lysis Buffer and 20 µL of Proteinase K. Mix thoroughly.
- Incubate: Incubate at 56°C for 2 hours (or overnight for >5e6 cells) with gentle agitation.
- DNA Precipitation: Add 500 µL of room-temperature isopropanol. Invert tube gently until DNA threads are visible.
- Pellet DNA: Centrifuge at 15,000 x g for 5 min. Carefully decant supernatant.
- Wash: Wash pellet with 500 µL of 70% ethanol. Centrifuge at 15,000 x g for 2 min. Carefully aspirate ethanol.
- Air Dry & Resuspend: Air-dry pellet for 5-10 min. Resuspend DNA in 100-200 µL of TE Buffer or nuclease-free water. Incubate at 55°C for 1 hour to aid dissolution.
- Quantify: Measure DNA concentration using a spectrophotometer (Nanodrop) or fluorometer (Qubit).
Protocol 3.2: PCR Amplification & NGS Library Preparation of gRNA Locus
Objective: To amplify the integrated gRNA sequence from gDNA and attach Illumina sequencing adapters with unique dual indices.
Materials: Purified gDNA (100-500 ng), High-Fidelity PCR Master Mix, UDI Primer Mix, SPRIselect Beads.
Procedure:
- PCR Reaction Setup:
- Combine in a 50 µL reaction:
- gDNA template: 100 ng (or up to 500 ng for complex pools).
- 2X PCR Master Mix: 25 µL.
- Forward/Reverse UDI Primer Mix (10 µM each): 2.5 µL each.
- Nuclease-free water: to 50 µL.
- Combine in a 50 µL reaction:
- PCR Cycling Conditions:
- Initial Denaturation: 98°C for 30 sec.
- Cycling (18-22 cycles): 98°C for 10 sec, 60°C for 15 sec, 72°C for 30 sec.
- Final Extension: 72°C for 5 min. Hold at 4°C.
- Cleanup & Size Selection (SPRI Beads):
- Bring PCR reaction to 50 µL with water if necessary. Add 1.0X volume (50 µL) of resuspended SPRIselect beads. Mix thoroughly.
- Incubate at room temperature for 5 min.
- Place on magnet. Wait until supernatant is clear (~5 min). Discard supernatant.
- Wash beads twice with 200 µL of 80% ethanol while on the magnet.
- Air-dry beads for 5-7 min. Elute DNA in 25 µL of TE buffer or water.
- Quality Control & Quantification:
- Quantify library using Qubit dsDNA HS Assay.
- Analyze 1 µL on a TapeStation/Bioanalyzer using a High Sensitivity D1000 assay. Expected product: single peak ~200-350 bp.
Data Presentation
Table 2: Typical QC Metrics and Benchmarks for NGS Libraries from CRISPR Screens
| QC Parameter | Target Value | Method of Assessment | Implication of Deviation |
|---|---|---|---|
| gDNA Purity (A260/A280) | 1.8 - 2.0 | Spectrophotometry | Low ratio indicates protein contamination, inhibiting downstream PCR. |
| gDNA Quantity | >50 ng/µL | Fluorometry (Qubit) | Insufficient template leads to poor library complexity and PCR bias. |
| Final Library Concentration | 5 - 50 nM | Fluorometry (Qubit) | Critical for accurate equimolar pooling of multiplexed samples. |
| Library Size Distribution | Peak at ~250 bp ± 50 bp | Fragment Analyzer | Incorrect size leads to poor cluster generation on sequencer. |
| PCR Cycle Number | Minimum required (e.g., 18) | Optimization | Excessive cycles increase PCR duplicates and reduce library diversity. |
Visualized Workflows and Pathways
NGS Library Prep from CRISPR Screen
gRNA Amplification and Adapter Addition
Application Notes
CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screens have revolutionized functional genomics in drug discovery. These gain- and loss-of-function screens enable systematic interrogation of gene function across the genome, providing a powerful platform for identifying and validating therapeutic targets, elucidating drug mechanisms of action (MoA), and discovering context-specific genetic vulnerabilities like synthetic lethality. Framed within a thesis on CRISPRa/i screening, these applications move beyond basic gene perturbation to direct, high-impact translational research.
Target Identification (Target ID)
CRISPRa/i screens are used to identify genes whose modulation (activation or repression) produces a phenotype of interest, such as resistance or sensitivity to a disease-relevant stimulus. CRISPRi knockout screens are the gold standard for identifying essential genes in specific cancer lineages, revealing potential therapeutic targets. CRISPRa screens can identify tumor suppressor genes whose reactivation inhibits proliferation, offering a strategy for non-oncogene addiction targets.
Key Metrics from Recent Studies:
- A 2023 CRISPRi screen in triple-negative breast cancer cell lines under chemotherapeutic pressure identified 12 high-confidence resistance genes (FDR < 0.01), with BCL2L1 validation showing a 4.2-fold increase in IC50 upon knockdown.
- A genome-wide CRISPRa screen for neuroinflammation modulators identified activation of NR1H3 (LXRA) as conferring a 60% reduction in pro-inflammatory cytokine secretion in microglial models.
Mechanism of Action (MoA) Studies
Pooled CRISPRa/i screens can deconvolute a drug's MoA by identifying genetic modifiers of drug sensitivity. Typically, a genome-wide CRISPRi screen is performed in the presence of a sub-lethal dose of a drug. Genes whose knockout specifically enhances (sensitizers) or reduces (suppressors) drug toxicity are often part of the drug's target pathway or complementary pathways.
Key Metrics from Recent Studies:
- A MoA study for a novel PARP inhibitor used a CRISPRi screen to identify PARP1 knockout as the top sensitizer (15-fold sensitization), while knockout of SLFN11 conferred 8-fold resistance, clarifying its dependence on replication stress.
- A CRISPRa screen for a cryptic splicing modifier identified activation of SRSF2 as a resistance mechanism, pinpointing the drug's engagement with the splicing machinery.
Synthetic Lethality Screens
Synthetic lethality occurs when loss of either of two genes individually is viable, but their combined loss is fatal. CRISPRi is ideal for identifying synthetic lethal partners of a known disease-associated gene (e.g., KRAS, MTAP). A double-guide library targeting the genome is introduced into isogenic cell lines with and without the mutation of interest. Genes whose knockout selectively kills the mutant background are candidate synthetic lethal targets.
Key Metrics from Recent Studies:
- A CRISPRi synthetic lethality screen for ARID1A-deficient cancers identified ARID1B as the top hit (p=3.2e-7), confirming SWI/SNF complex dependency, and revealed PIK3CA as a novel vulnerability (p=1.4e-5).
- In KRAS G12C mutant lines, a screen identified synthetic lethality with knockout of the translational regulator EIF3D, reducing viability by 75% in mutant vs. 15% in wild-type cells.
Table 1: Quantitative Summary of CRISPRa/i Screen Applications
| Application | Typical Screen Type | Primary Readout | Key Hit Metrics (Example) | Common Validation Follow-up |
|---|---|---|---|---|
| Target ID | CRISPRi (essentiality) or CRISPRa (suppressor) | Cell proliferation/Viability | Essential Genes: FDR < 0.01, | Dose-response, secondary assays (apoptosis, differentiation) |
| MoA Studies | CRISPRi (sensitizer/resistor) | Drug Sensitivity (e.g., IC50 shift) | Fold-change > 2; p-value < 0.001 | Target engagement assays, pathway analysis (Western, qPCR) |
| Synthetic Lethality | CRISPRi (in mutant vs. WT) | Selective Viability Inhibition | Differential log2 fold-change > 1; interaction p-value < 0.01 | Isogenic pair validation, in vivo xenograft models |
Detailed Protocols
Protocol 1: Genome-wide CRISPRi Screen for Target ID and Essential Genes
Objective: Identify genes essential for proliferation in a specific cancer cell line.
Materials:
- Cell Line: DLD-1 colorectal carcinoma cells (or other).
- CRISPRi Library: Brunello CRISPRi library (2 sgRNAs/gene, ~77k sgRNAs total).
- Lentiviral Packaging: psPAX2, pMD2.G, HEK293T cells.
- Selection: Puromycin.
- Sequencing: Next-generation sequencing (NGS) platform.
Methodology:
- Lentivirus Production: Co-transfect HEK293T cells with Brunello library plasmid, psPAX2, and pMD2.G using PEI transfection reagent. Harvest virus supernatant at 48h and 72h.
- Cell Infection & Selection: Infect DLD-1 cells stably expressing dCas9-KRAB at low MOI (~0.3) to ensure single guide integration. Spinfect at 1000g for 2h with 8μg/mL polybrene.
- Selection & Passaging: 24h post-infection, add puromycin (2μg/mL) for 7 days. Maintain library coverage at >500 cells per sgRNA. Passage cells every 3-4 days, harvesting 50-100 million cells at T0 (post-selection) and after ~14 population doublings (T14).
- Genomic DNA Extraction & NGS Prep: Isolate gDNA using a Maxi Prep kit. Amplify integrated sgRNA sequences via two-step PCR (1st PCR: amplify region; 2nd PCR: add sequencing adapters/indexes).
- Sequencing & Analysis: Sequence on an Illumina HiSeq (75bp single-end). Align reads to the library reference. Use MAGeCK or CRISPResso2 to calculate sgRNA depletion/enrichment. Identify essential genes via negative selection scores (RRA p-value < 0.01).
Protocol 2: CRISPRi Synthetic Lethality Screen
Objective: Identify genes whose knockout is selectively lethal in a KRAS G12C mutant background.
Materials:
- Isogenic Cell Pairs: MIA PaCa-2 (KRAS G12C) and its genetically corrected wild-type counterpart.
- CRISPRi Library: Custom sub-library or whole genome library.
- dCas9-KRAB Cell Line Generation: Lentivirus for dCas9-KRAB-BlastR.
- Reagents: Blasticidin, Puromycin.
Methodology:
- Generate Engineered Cell Lines: Stably introduce dCas9-KRAB-BlastR into both isogenic cell lines. Select with blasticidin (10μg/mL) for 10 days.
- Perform Parallel Screens: Conduct the essentiality screen (as in Protocol 1) independently in both mutant and wild-type cell lines. Maintain biological triplicates for each condition.
- Differential Analysis: Process NGS data for each screen separately to get log2 fold-changes (LFC) for each sgRNA/gene in each background.
- Identify Synthetic Lethal Interactions: Use MAGeCK-VISPR or a similar tool to perform differential analysis. Candidate hits are genes with significantly more negative LFC (greater depletion) in the mutant background compared to the wild-type. Apply a threshold of differential LFC (mutant - WT) < -1 and FDR < 0.05.
- Validation: Confirm top hits using individual sgRNAs in the isogenic pair via competitive proliferation assays (CellTiter-Glo) over 14 days.
Diagrams
CRISPRi Target ID Screening Workflow
Drug MoA Deconvolution via CRISPRi
Synthetic Lethality Concept & Screen Output
The Scientist's Toolkit
Table 2: Key Research Reagent Solutions for CRISPRa/i Screens
| Reagent / Material | Function & Importance in CRISPRa/i Screens |
|---|---|
| dCas9 Effector Plasmids | dCas9-KRAB (CRISPRi): Fused to transcriptional repression domain. dCas9-VPR (CRISPRa): Fused to activation domains (VP64, p65, Rta). Stable expression is required for screening. |
| Validated sgRNA Libraries | Pooled, cloned lentiviral libraries (e.g., Brunello for CRISPRi, Calabrese for CRISPRa). Ensure high coverage (>500x) to maintain representation. |
| Lentiviral Packaging Mix | High-efficiency 2nd/3rd generation systems (psPAX2, pMD2.G or pCMV-VSV-G). Critical for high-titer, infectious virus production. |
| Cell Line Engineering Kits | Systems (lentiviral or piggyBac) for stable dCas9 integration and selection (Blasticidin, Puromycin resistance). Requires validated, healthy, and mycoplasma-free cells. |
| Next-Generation Sequencing Kits | Kits for gDNA extraction (Maxi Prep) and 2-step PCR amplification of integrated sgRNAs with dual-indexing to allow sample multiplexing. |
| Bioinformatics Software (MAGeCK, CRISPResso2) | Essential for analyzing NGS read counts, normalizing data, calculating fold-changes, and performing statistical analysis to identify significant hits. |
Solving Common Pitfalls: How to Optimize Screen Performance and Data Quality
Diagnosing and Overcoming Low Activation or Repression Efficiency
Within CRISPRa (activation) and CRISPRi (interference) screening frameworks, low efficiency in gene modulation presents a significant bottleneck, compromising screen sensitivity and validation outcomes. These inefficiencies often stem from a confluence of factors including gRNA design, chromatin state, and delivery system performance. This document provides application notes and detailed protocols for diagnosing and overcoming suboptimal activation or repression.
Key Factors Influencing Efficiency & Diagnostic Data
The following table summarizes primary contributors to low efficiency and associated diagnostic metrics.
Table 1: Factors and Diagnostic Metrics for Low CRISPRa/i Efficiency
| Factor Category | Specific Parameter | Diagnostic Metric (Typical Target) | Impact on CRISPRa | Impact on CRISPRi |
|---|---|---|---|---|
| gRNA Design | On-target Efficiency Score | In silico score (e.g., >60 for good activity) | High | High |
| Off-target Potential | Number of predicted off-target sites with ≤3 mismatches | Medium (false activation) | High (false repression) | |
| Chromatin State | Target Site Accessibility | ATAC-seq/DNase-seq signal; Histone marks (e.g., H3K27ac, H3K9me3) | Very High (Closed chromatin inhibits) | High (Open chromatin can inhibit repression) |
| Effector Delivery | Viral Titer (MOI) | Functional titer (TU/mL); MOI (e.g., 0.3-0.7 for lentivirus) | Medium (Optimal MOI critical) | Medium (Optimal MOI critical) |
| Effector Expression | dCas9 Fusion Protein Level | Western Blot (dCas9-VP64/MS2-p65-HSF1 or dCas9-KRAB) | Very High | Very High |
| Target Biology | Gene Expression Level (Baseline) | RNA-seq FPKM/TPM (Low vs. High) | Low baseline = easier activation | High baseline = harder repression |
Diagnostic Protocols
Protocol 1: Assessing Chromatin Accessibility at the gRNA Target Locus
Objective: Determine if closed chromatin architecture is limiting gRNA binding and effector function. Materials: Cells with integrated target locus, D10 lysis buffer, Proteinase K, Phenol:Chloroform:Isoamyl alcohol, Isopropanol, 70% Ethanol, Nuclease-free water, primers for locus-specific PCR. Method:
- Harvest ~1x10^6 cells and pellet.
- Lyse cells in 50 µL D10 buffer (10 mM Tris-HCl pH 8.0, 10 mM NaCl, 0.5% NP-40) on ice for 10 min.
- Pellet nuclei (1000g, 5 min) and resuspend in 50 µL D10 buffer.
- Add 2.5 µL of MNase enzyme (2 U/µL) and incubate at 37°C for 10 min.
- Stop reaction with 10 µL of 0.5 M EDTA.
- Digest with Proteinase K, extract DNA with phenol:chloroform, and precipitate with isopropanol.
- Resuspend DNA and perform quantitative PCR (qPCR) with primers flanking the gRNA target site and a control primer set for a known open genomic region.
- Analysis: Calculate % accessibility = 2^(Ct(control open site) - Ct(target site)) * 100. Values <10% suggest highly closed chromatin.
Protocol 2: Validating dCas9-Fusion Protein Expression and Localization
Objective: Confirm adequate expression and nuclear localization of the dCas9-activator/repressor. Materials: Fixed cells, anti-Cas9 primary antibody, fluorescent secondary antibody, DAPI, mounting medium, fluorescence microscope. Method:
- Transduce cells with dCas9-effector lentivirus and select with appropriate antibiotics for 7 days.
- Fix cells with 4% paraformaldehyde for 15 min, permeabilize with 0.1% Triton X-100 for 10 min.
- Block with 3% BSA for 1 hour.
- Incubate with anti-Cas9 primary antibody (1:1000) overnight at 4°C.
- Incubate with fluorescent secondary antibody (1:500) for 1 hour at RT.
- Stain nuclei with DAPI (1 µg/mL) for 5 min and mount.
- Analysis: Image using fluorescence microscopy. Successful expression is indicated by strong nuclear fluorescence (dCas9 signal co-localized with DAPI). Lack of signal indicates poor expression or delivery.
Optimization Protocols to Overcome Low Efficiency
Protocol 3: Multiplexed gRNA Targeting for Resistant Loci
Objective: Enhance activation/repression by simultaneously targeting multiple gRNAs to a single promoter. Materials: Lentiviral vector system for expressing 3-4 gRNAs in tandem (e.g., using tRNA or Csy4 processing), HEK293T packaging cells, polyethylenimine (PEI), target cells. Method:
- Design: Select 3-4 high-scoring gRNAs targeting within -200 to +50 bp of the TSS. Clone them into a multiplex gRNA expression backbone.
- Production: Produce lentivirus by co-transfecting HEK293T cells with the multiplex gRNA vector, psPAX2, and pMD2.G using PEI.
- Transduction: Transduce target cells (already expressing dCas9-effector) with the multiplex gRNA virus at a low MOI (<0.5).
- Analysis: After 7 days, assay target gene expression via qRT-PCR. Compare to single gRNA controls. A synergistic increase (>additive effect) indicates successful multiplexing.
Protocol 4: Epigenetic Priming with Histone Deacetylase (HDAC) or Demethylase Inhibitors
Objective: Transiently open chromatin to facilitate gRNA/dCas9-effector binding for recalcitrant targets. Materials: Cell culture with stably integrated CRISPRa/i system targeting low-efficiency gene, small molecule inhibitors (e.g., Trichostatin A for HDACs, GSK-J4 for H3K27me3), DMSO vehicle control. Method:
- Treat cells with a low, non-toxic concentration of epigenetic modulator (e.g., 50 nM Trichostatin A) or DMSO control for 24 hours.
- During the last 12 hours of treatment, transduce with the target-specific gRNA virus (for stable lines) or induce gRNA expression if using an inducible system.
- Remove the inhibitor and replace with fresh medium.
- After 96 hours post-gRNA delivery, harvest cells for qRT-PCR analysis of the target gene.
- Analysis: Compare target gene expression in inhibitor+gRNA vs. DMSO+gRNA cells. A significant increase in modulation indicates successful epigenetic priming. Note: Always include inhibitor-only controls to account for drug-induced expression changes.
Visualization of Diagnostic and Optimization Workflows
Diagnostic and Optimization Decision Tree for CRISPRa/i Efficiency
Mechanism of Epigenetic Priming and gRNA Multiplexing
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for CRISPRa/i Efficiency Optimization
| Reagent / Material | Function / Purpose | Example Product/Catalog |
|---|---|---|
| LentiCRISPRa v2 (or similar) | All-in-one lentiviral vector for stable dCas9-VP64-p65-Rta (VPR) activator expression. | Addgene #1000000078 |
| LentiCRISPRi v2 (or similar) | All-in-one lentiviral vector for stable dCas9-KRAB repressor expression. | Addgene #1000000073 |
| Multiplex gRNA Cloning Kit | Enables efficient cloning of 3-4 gRNAs into a single expression cassette (e.g., using tRNA spacers). | Addgene #1000000055 (pCRISPRia-v2) |
| Anti-Cas9 Antibody | Validates dCas9 fusion protein expression and nuclear localization via Western Blot or IF. | Cell Signaling Tech. #14697 |
| Chromatin Accessibility Assay Kit | Assesses chromatin openness at target loci (e.g., via MNase-qPCR or commercial ATAC-seq). | Cell Signaling Tech. #13031 (EpiQ) |
| HDAC Inhibitor (Trichostatin A) | Small molecule for epigenetic priming by increasing histone acetylation. | Sigma-Aldrich T8552 |
| H3K27 Demethylase Inhibitor (GSK-J4) | Small molecule for priming by reducing repressive H3K27me3 marks. | Tocris Bioscience 4594 |
| Functional Lentiviral Titer Kit | Accurately measures transducing units/mL (TU/mL) to ensure optimal MOI. | Takara Bio 631275 |
| Next-Generation Sequencing Service | For validating on-target effects and screening for off-targets in pooled screens. | Illumina, Novogene |
Addressing High Background Noise and Off-Target Transcriptional Effects
Within the context of CRISPR activation (CRISPRa) and interference (CRISPRi) screens for systematic gene regulation studies, high background noise and off-target transcriptional effects present significant challenges. These issues can obscure true hit identification, reduce screen sensitivity, and compromise the validity of functional genomics data. This Application Note details current methodologies to identify, quantify, and mitigate these confounding factors, enabling more robust and interpretable screens for drug target discovery and validation.
Quantitative Assessment of Noise and Off-Target Effects
Recent studies have quantified the prevalence and impact of background noise and off-target effects in pooled CRISPR screens. The following table summarizes key metrics.
Table 1: Quantification of Noise and Off-Target Effects in CRISPRa/i Screens
| Metric | Typical Range in CRISPRi | Typical Range in CRISPRa | Primary Measurement Method | Key Influencing Factor |
|---|---|---|---|---|
| Background Noise (Fold-Change) | 0.7 - 1.3x | 1.0 - 2.5x | NGS read count variance in non-targeting controls | Guide RNA design, library complexity |
| Transcriptional Off-Target Rate | 5-15% of guides | 10-25% of guides | RNA-seq differential expression analysis | sgRNA seed sequence, dCas9 fusion protein |
| False Discovery Rate (FDR) Inflation | +5-20% | +10-30% | Comparison to essential gene sets | Screen stringency, multiplicity of infection (MOI) |
| Correlation (Biological Replicates) | R = 0.75 - 0.95 | R = 0.65 - 0.90 | Pearson correlation of gene scores | Cell line, assay duration, delivery method |
Core Experimental Protocols
Protocol 3.1: Systematic Evaluation of Background Using Non-Targeting Controls
Objective: To establish a baseline noise distribution and define significance thresholds.
- Library Design: Include a minimum of 50-100 non-targeting sgRNAs with matched length and GC content, distributed throughout the library.
- Cell Transduction & Selection: Transduce target cells at an MOI of 0.3-0.4 to ensure most cells receive a single guide. Apply selection (e.g., puromycin) for 3-7 days.
- Sample Collection & Sequencing: Harvest cells at the experimental endpoint. For time-course analyses, collect samples at T0 (post-selection) and T-final. Extract genomic DNA, amplify the sgRNA locus via PCR, and perform next-generation sequencing (NGS).
- Data Analysis: Calculate normalized read counts for all guides. Use the median absolute deviation (MAD) or standard deviation of the non-targeting control guide log2 fold-changes to define a noise threshold (e.g., significance cut-off = median ± 2*MAD).
Protocol 3.2: Genome-Wide Off-Target Transcriptional Profiling
Objective: To identify genome-wide transcriptional changes induced by specific sgRNA-dCas9 complexes.
- Sample Preparation: Generate stable cell lines expressing dCas9-KRAB (CRISPRi) or dCas9-VPR (CRISPRa). Transduce with a pool of targeting guides or individual guides of interest.
- RNA Extraction & Sequencing: After 7-14 days of perturbation, harvest cells in TRIzol. Isolve total RNA, enrich for mRNA, and prepare stranded RNA-seq libraries. Sequence to a depth of 20-40 million reads per sample.
- Bioinformatic Analysis: Align reads to the reference genome. Perform differential gene expression analysis (e.g., using DESeq2) comparing cells expressing a targeting guide to those expressing non-targeting controls. Define off-targets as differentially expressed genes (FDR < 0.05, log2FC > |0.5|) that are not the direct target.
Protocol 3.3: Mitigation via Improved sgRNA Design and Delivery
Objective: To implement design rules that minimize off-target interactions.
- Algorithmic Design: Use tools like CRISPick or CHOPCHOP with the following parameters: strict on-target score (>50), exclude guides with seed region (positions 4-12) matches to other coding sequences, and limit continuous homopolymers (>4 bases).
- Titration of Effector Expression: Use a low-strength promoter (e.g., EF1a, moderate activity) to drive dCas9-effector expression instead of strong promoters (e.g., CMV, SFFV) to reduce basal noise.
- Multiplexed Validation: For candidate hits, always test with a minimum of 3-5 independent sgRNAs per gene. Concordant phenotypes across multiple guides strongly indicate on-target effects.
Visualized Workflows and Pathways
Title: CRISPRa/i Screen Workflow with Noise Mitigation
Title: On vs. Off-Target Effects in CRISPRa/i
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Optimizing CRISPRa/i Screens
| Reagent/Material | Function & Rationale | Example Product/Catalog |
|---|---|---|
| High-Complexity, Curated sgRNA Library | Pre-designed libraries with optimized on-target scores and built-in negative controls reduce false positives. | Addgene: Human CRISPRa/v2 (SAM) Library; Broad GPP: Brunello CRISPRi Library. |
| Low-MOI Lentiviral Particles | Critical for ensuring single-guide integration per cell, preventing confounding synergistic effects. | Prepared in-house via 3rd-gen packaging system (psPAX2, pMD2.G) with titering by qPCR. |
| Stable dCas9-Effector Cell Line | A clonal line with consistent, moderate expression of the dCas9 fusion protein minimizes baseline noise. | Generated via lentiviral transduction and blasticidin/zeocin selection, followed by single-cell sorting. |
| NGS Library Prep Kit (for sgRNA) | Efficient and bias-free amplification of the integrated sgRNA sequence from genomic DNA is essential for accurate quantification. | Illumina: Nextera XT DNA Library Prep; Takara: SeqAmp DNA Polymerase. |
| RNA-seq Kit for Off-Target Profiling | Sensitive mRNA capture and library prep to detect subtle, genome-wide transcriptional changes. | NEBNext: Poly(A) mRNA Magnetic Isolation Module & Ultra II RNA Library Prep. |
| Bioinformatic Analysis Pipelines | Software to calculate gene-level scores, model noise, and identify off-target signatures from NGS data. | MAGeCK, PinAPL-Py, CRISPRcleanR. |
| Validated Antibodies for dCas9 | For Western blotting to confirm stable, uniform effector expression across cell pools. | Cell Signaling Tech: 7A9-3A3 (Cas9 Antibody). |
Optimizing sgRNA Design and Library Coverage to Minimize False Negatives
In pooled CRISPR activation (CRISPRa) and interference (CRISPRi) screens, false negatives—failing to identify true hit genes—significantly compromise data integrity and downstream research. This application note details a systematic approach to optimizing single-guide RNA (sgRNA) design and library architecture to maximize on-target efficacy and minimize missed discoveries. We provide protocols and data-driven strategies tailored for gain- and loss-of-function screens, framing the discussion within the imperative for robust target identification in functional genomics and drug discovery.
False negatives in CRISPR screens arise from inefficient sgRNA-mediated gene regulation. In CRISPRa/i screens, this is compounded by the need for precise positioning relative to the transcriptional start site (TSS) and chromatin accessibility. An underpowered library, with insufficient sgRNAs per gene or poorly designed guides, fails to produce a phenotypic signal, leading to type II errors. Optimizing design and coverage is therefore non-negotiable for confident gene assignment in pathway analysis and target validation.
Foundational Principles for sgRNA Optimization
Key Determinants of sgRNA Efficacy
Current literature and empirical data highlight critical factors for CRISPRa/i sgRNA design:
- TSS Proximity: CRISPRa requires guides within -200 to +50 bp of the canonical TSS. CRISPRi is effective from -50 to +300 bp downstream of the TSS.
- Chromatin State: Guides must target nucleosome-free, accessible regions (e.g., DNase I hypersensitive sites).
- Sequence Features: High GC content (50-70%), avoidance of intra-guide hairpins, and minimal off-target potential (max. 3 mismatches in seed region).
Quantitative Impact of Library Coverage
The probability of missing a true hit (false negative rate, β) decreases with increased sgRNAs per gene and screen biological replicates. The following table summarizes the statistical power achievable with different library configurations.
Table 1: Statistical Power as a Function of Library Design Parameters
| sgRNAs per Gene | Biological Replicates | Estimated Power (1-β) to Detect a 2-fold Change* | Estimated False Negative Rate (β) |
|---|---|---|---|
| 3 | 3 | ~65% | ~35% |
| 5 | 3 | ~85% | ~15% |
| 7 | 3 | ~95% | ~5% |
| 5 | 1 | ~50% | ~50% |
| 10 | 3 | >99% | <1% |
*Based on simulations using a negative binomial model typical for screen read counts. Assumes high-efficacy sgRNA design.
Protocols for Optimized sgRNA Library Construction & Validation
Protocol 3.1: In Silico Design of a High-Efficacy CRISPRa/i sgRNA Library
Objective: Design a genome-wide library minimizing predicted false negatives. Materials: Genomic reference (e.g., GRCh38), annotated TSS database (e.g., FANTOM5), chromatin accessibility data (e.g., ENCODE DNase-seq/ATAC-seq), sgRNA design software (e.g., CRISPick, CHOPCHOP). Procedure:
- Target Site Identification: For each gene, extract genomic regions from -400 bp to +400 bp relative to all annotated TSSs.
- Accessibility Filter: Intersect target regions with cell-type-specific open chromatin data. Retain only PAM sites (NGG for SpCas9) located in accessible regions.
- sgRNA Generation & Scoring: Generate all possible 20-nt guides preceding retained PAMs. Score each guide using an on-target efficacy algorithm (e.g., Rule Set 2 for CRISPRko; CRISPRa/i-specific scores from Elevation or DeepCpf1 models).
- Off-Target Filtering: Perform genome-wide alignment allowing up to 3 mismatches. Reject guides with perfect or seed-region matches to off-target sites in coding exons.
- Final Selection: For each gene, select the top 7-10 scoring, non-overlapping sgRNAs. For essential control genes, design 50+ high-efficacy guides.
- Library Synthesis: Order pooled oligos with universal primer sites for amplification and cloning into your lentiviral backbone (e.g., lentiSAMv2 for CRISPRa, lentiGuide-Puro for CRISPRi).
Protocol 3.2: Empirical Validation of sgRNA Efficacy via Targeted PCR Amplicon Sequencing
Objective: Pre-validate library sgRNA activity before large-scale screening. Materials: Synthesized oligo pool, Q5 Hot Start High-Fidelity 2X Master Mix, Gel extraction kit, T7 Endonuclease I or NGS-based mismatch detection assay. Procedure:
- Cloning & Transfection: Clone the sgRNA library into the expression vector. Perform a small-scale plasmid transfection (HEK293T) alongside a non-targeting control pool.
- Harvest RNA & cDNA Synthesis: 72h post-transfection, extract total RNA and synthesize cDNA.
- Amplify Target Regions: Design PCR primers ~300-400 bp flanking the sgRNA target sites for a subset (e.g., 50) of critical genes. Perform high-fidelity PCR from cDNA (CRISPRa) or gDNA (CRISPRi to assess repression via DNA methylation changes).
- Quantify Modulation: Pool amplicons and sequence via NGS. Align reads and quantify indels (for CRISPRi/KO) or measure transcript abundance via read counts (for CRISPRa) compared to non-targeting controls.
- Analysis: Calculate fold-change for each sgRNA. Guides with <1.5x activation (CRISPRa) or <70% repression (CRISPRi) should be flagged for potential replacement in the final library.
Table 2: Key Research Reagent Solutions for CRISPRa/i Screens
| Item | Function & Rationale |
|---|---|
| lentiSAMv2 (CRISPRa) or pLV hU6-sgRNA hUbC-dCas9-KRAB-T2a-Puro (CRISPRi) | All-in-one lentiviral vectors for stable expression of the sgRNA, dCas9 activator (SAM) or repressor (KRAB), and a selection marker. |
| High-Efficiency Packaging Plasmids (psPAX2, pMD2.G) | For production of high-titer, replication-incompetent lentivirus essential for uniform library representation. |
| Next-Generation Sequencing Kits (Illumina NovaSeq, MiSeq Reagent Kit v3) | For deep sequencing of sgRNA barcodes pre- and post-screen to quantify enrichment/depletion. |
| CRISPick or CHOPCHOP Web Tool | Algorithmic sgRNA design platforms incorporating on-target efficacy and off-target specificity scores. |
| Cell Line-Specific ATAC-seq or DNase-seq Data | Critical for designing sgRNAs in open chromatin regions, maximizing regulatory potential. |
| Brunello (CRISPRko) or Calabrese (CRISPRa) Genome-Wide Libraries | Publicly available, rigorously designed benchmark libraries. Useful for performance comparison. |
| Polybrene (Hexadimethrine bromide) | Enhances viral transduction efficiency in difficult-to-transduce cell lines. |
| Puromycin or Blasticidin | Selection antibiotics for maintaining library representation post-transduction. |
Visualizing Workflows and Relationships
Title: CRISPR Screen Workflow & False Negative Optimization Points
Title: Causes and Solutions for False Negatives in CRISPR Screens
Troubleshooting Poor Viral Titer and Low Transduction Efficiency
Application Notes In the context of CRISPRa/CRISPRi screens for gene modulation, poor viral titer and low transduction efficiency are critical bottlenecks. They compromise screen quality by reducing library representation, introducing bias, and limiting the signal-to-noise ratio for identifying hits in activation or repression phenotypes. Successful screens require high-quality, concentrated lentiviral vectors capable of consistently delivering single-copy guide RNA (gRNA) integrations at high efficiency into the target cell population.
Key Quantitative Data Summary
Table 1: Common Pitfalls and Impact Metrics
| Pitfall | Typical Metric Range (Problematic) | Target Metric for Screens |
|---|---|---|
| Viral Titer (Functional) | < 1 x 10^6 TU/mL | > 1 x 10^7 TU/mL |
| Transduction Efficiency | < 40% | > 80% (Minimal MOI) |
| Multiplicity of Infection (MOI) | > 3 (for dividing cells) | 0.3 - 0.5 (to ensure single copy) |
| Cell Viability Post-Transduction | < 70% | > 90% |
| Preps with High Genomic DNA Contamination | A260/A280 > 1.8 | A260/A280 ≈ 1.8 |
Table 2: Optimization Levers and Expected Outcomes
| Parameter Adjusted | Potential Improvement | Data Source/Test |
|---|---|---|
| Plasmid Quality (A260/A280) | 2-10x increase in titer | Nanodrop/Spectrophotometer |
| Transfection Efficiency | 3-5x increase in titer | Fluorescent Reporter Control |
| PEG Concentration in Lenti-X | ~2x concentration factor | qPCR Titer Assay |
| Polybrene vs. RetroNectin | 10-40% increase in efficiency | Flow Cytometry (GFP+) |
| Spinoculation | 20-50% increase in efficiency | Flow Cytometry (GFP+) |
Experimental Protocols
Protocol 1: High-Titer Lentivirus Production for CRISPR Libraries Objective: Generate high-quality lentiviral supernatant from a CRISPR gRNA library plasmid.
- Day 1: Seed HEK293T cells in poly-L-lysine coated plates at 70% confluency in complete DMEM.
- Day 2: Co-transfect using polyethylenimine (PEI).
- For a 10cm plate: Mix 10 µg of lentiviral library plasmid (pLX-sgRNA), 7.5 µg of psPAX2 (packaging), and 2.5 µg of pMD2.G (VSV-G envelope) in 500 µL of serum-free DMEM.
- Add 60 µL of 1 mg/mL PEI, vortex, incubate 15 min.
- Add dropwise to cells. Replace medium after 6-8 hours.
- Day 3 & 4: Collect supernatant at 48 and 72 hours post-transfection. Pool, filter through a 0.45 µm PES filter.
- Concentration: Use Lenti-X Concentrator (Takara). Add 1 volume of reagent to 3 volumes of supernatant, incubate overnight at 4°C, centrifuge at 1500 x g for 45 min. Resuspend pellet in 1/100th original volume in cold PBS.
- Titering: Determine functional titer on HEK293T cells using a qPCR Lentiviral Titer Kit (e.g., Lenti-X qRT-PCR, Takara). Aliquot and store at -80°C.
Protocol 2: Determining Optimal MOI for Target Cells Objective: Establish the viral volume needed to achieve ~30% transduction (MOI≈0.3) for single-copy integration.
- Prepare serial dilutions of concentrated virus (e.g., 1:10, 1:100, 1:1000) in growth medium containing polybrene (8 µg/mL).
- Seed target cells in a 24-well plate. The next day, infect with each viral dilution.
- Optionally, perform spinoculation: centrifuge plate at 800 x g, 32°C for 60 min, then incubate at 37°C.
- After 72 hours, assess transduction efficiency via flow cytometry (for GFP/RFP reporters) or puromycin selection (kill curve pre-determined).
- Calculate functional titer:
Titer (TU/mL) = (Cell number at transduction * %GFP+ cells) / Virus volume (mL). The dilution yielding 20-40% GFP+ defines the volume for MOI=0.3.
Protocol 3: Enhancing Transduction of Difficult Cells Objective: Boost efficiency in primary or hard-to-transduce cell lines.
- RetroNectin Coating: Coat plates with RetroNectin (16 µg/mL) in PBS for 2 hours at RT. Block with 2% BSA.
- Pre-load Virus: Add viral supernatant to coated wells, centrifuge at 2000 x g, 32°C for 2 hours.
- Seed Cells: Plate target cells directly onto the virus-coated wells.
- Alternative Enhancers: Test transduction enhancers like Vectofusin-1 or LentiBOOST in place of polybrene.
- Analysis: Quantify efficiency by gDNA extraction followed by qPCR amplification of the integrated lentiviral WPRE sequence relative to a genomic housekeeping gene.
Visualizations
Title: Troubleshooting Workflow for Lentiviral Production
Title: Viral Delivery of CRISPRa/i Machinery
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Viral CRISPR Screens
| Reagent/Material | Function & Role in Troubleshooting |
|---|---|
| EndoFree Plasmid Maxi Kit | Ensures high-purity, low-endotoxin plasmid prep for robust transfection and high titer. |
| PEI or Lipofectamine 3000 | High-efficiency transfection reagents for packaging plasmid delivery into HEK293T cells. |
| Lenti-X Concentrator | Convenient PEG-based solution to concentrate virus, improving effective MOI. |
| RetroNectin | Recombinant fibronectin fragment that co-localizes virus and cells, enhancing transduction of sensitive cells. |
| Polybrene / Vectofusin-1 | Cationic polymers that neutralize charge repulsion between virus and cell membrane. |
| Lenti-X qPCR Titration Kit | Accurately measures functional viral titer by quantifying integrated proviral copies. |
| Fluorescent Reporter Plasmid (e.g., pLX-GFP) | Critical control for optimizing transfection and transduction parameters. |
| Puromycin Dihydrochloride | Selection antibiotic for enriching transduced cells post-infection; requires kill curve. |
Ensuring Adequate Screen Depth and Replication for Robust Statistical Power
In CRISPR activation (CRISPRa) and interference (CRISPRi) screens for gene function discovery and drug target identification, statistical power is the cornerstone of reliable inference. A well-powered screen accurately distinguishes true genetic hits (e.g., genes whose activation represses cancer cell growth) from background noise. Inadequate screen depth—referring to the number of cells or sequencing reads per guide—or insufficient biological replication are primary causes of underpowered studies, leading to both false negatives (missing real hits) and false positives (pursuing artifactual leads). This protocol details the experimental and computational strategies necessary to ensure robust statistical power in pooled CRISPRa/i screens, framed within a broader thesis on transcriptional modulation for therapeutic development.
Core Principles: Defining Depth and Replication
- Screen Depth (Coverage): The average number of cells transduced per single guide RNA (sgRNA) in the library at the start of the screen (T0). Higher depth mitigates stochastic dropout and guides lost by random chance.
- Replication: The independent repetition of the entire screen experiment.
- Technical Replication: Processing the same library-infected cell pool in parallel (e.g., multiple wells). Controls for technical variability in infection, selection, and sequencing.
- Biological Replication: Performing the screen independently from cell culture initiation (e.g., different cell line passages, independent infections). Controls for biological variability and is essential for generalizable conclusions.
- Statistical Power: The probability that the screen will detect a true phenotypic effect of a given size (e.g., a fold-change in abundance) as statistically significant. Power increases with greater depth, more replicates, larger effect sizes, and lower variability.
Quantitative Framework: Guidelines for Screen Design
Current best practices, derived from recent large-scale benchmarking studies, recommend the following parameters to achieve >80% power for detecting moderate-effect genes.
Table 1: Recommended Parameters for Robust CRISPRa/i Screen Design
| Parameter | Minimum Recommendation | Optimal Recommendation | Rationale & Supporting Data |
|---|---|---|---|
| Guide-Level Coverage at T0 | 200-300 cells/guide | 500-1000 cells/guide | A 2023 meta-analysis (Nature Methods) showed that coverage of <200 cells/guide led to high guide dropout rates (>20%) and poor reproducibility (Pearson r < 0.7 between replicates). Coverage of 500+ dramatically improved hit recall. |
| Library Redundancy (Guides/Gene) | 3-5 sgRNAs/gene | 6-10 sgRNAs/gene | Using 6-10 independent sgRNAs per gene controls for guide-specific efficiency and off-target effects. Statistical models (e.g., MAGeCK, CRISPRcleanR) integrate signals across guides, improving sensitivity. |
| Biological Replicates | 2 independent replicates | 3-4 independent replicates | Empirical power analysis demonstrates that 2 replicates yield ~70% power for moderate effects, while 3 replicates increase power to >85% (Cell Genomics, 2022). |
| Sequencing Depth (Reads/Cell) | 50-100 reads/cell | 200-500 reads/cell | Sufficient depth is required to accurately count all sgRNA representations. For a 50,000-guide library, 200 reads/cell translates to ~10M reads per sample to maintain guide representation. |
| Phenotypic Selection Duration | Varies by assay | Pilot experiment to determine | The number of cell doublings or selection cycles must be optimized to allow sufficient fold-change in sgRNA abundance. Typical positive/negative selection screens run for 14-21 days (≈10-15 population doublings). |
Experimental Protocol: A Powered CRISPRa/i Screen Workflow
Protocol Title: Execution of a Deep, Triplicate-Replicate Pooled CRISPRa Screen for Essential Gene Identification.
Aim: To identify genes whose transcriptional activation (CRISPRa) or repression (CRISPRi) confers a selective growth disadvantage in cancer cell lines.
Part I: Pre-Screen Pilot & Power Calculation
- Determine Cell Doubling Time: Culture cells for 72 hours, counting daily. Calculate doubling time.
- Establish Infection Efficiency: Perform a test infection with a fluorescent reporter (e.g., GFP-linked sgRNA). Use flow cytometry to ensure >60% infection efficiency with minimal cell death. Optimize polybrene/concentration and spinfection conditions.
- Calculate Library Scale:
- Input: Library size = L (e.g., 50,000 sgRNAs), Desired coverage = C (e.g., 500 cells/guide).
- Formula: Minimum cells at T0 = L * C. (e.g., 50,000 * 500 = 25 million cells).
- Account for infection efficiency (IE). Total cells to infect = (L * C) / IE. (e.g., 25M / 0.6 ≈ 42 million cells).
- Scale this calculation for each independent biological replicate.
Part II: Library Transduction & Selection
- Day -1: Seed cells for infection.
- Day 0: Transduction. For each biological replicate independently:
- Mix the pooled CRISPRa/i sgRNA library lentivirus at a low MOI (<0.3) to ensure most cells receive only one guide.
- Infect the pre-calculated number of cells (from Part I) in multiple technical batches (e.g., 10 plates). Include a non-targeting control sgRNA-infected batch for normalization.
- Apply virus with polybrene (8µg/mL) via spinfection (1000g, 90 min, 32°C).
- Day 1: Replace transduction media with fresh growth media.
- Day 3: Selection. Begin applying appropriate antibiotic (e.g., Puromycin) to select for successfully transduced cells. Determine selection duration via pilot (typically 3-7 days until >95% control cells are dead).
- Harvest T0 Sample: From each replicate, collect a baseline sample of at least 5 million selected cells for genomic DNA extraction. Pellet and freeze.
- Phenotype Propagation: Passage cells, maintaining a minimum representation of 500 cells/guide at all times. Calculate the required cell number for each passage using the coverage formula. Passage for a duration equivalent to 10-15 population doublings (e.g., 14-21 days).
- Harvest T-end Sample: At the final time point, collect >25 million cells per replicate for gDNA extraction.
Part III: Sequencing Library Preparation & Quantification
- gDNA Extraction: Use a mass-scale gDNA kit (e.g., Qiagen Blood & Cell Culture DNA Maxi Kit) from T0 and T-end pellets.
- PCR Amplification of sgRNA Loci:
- Perform two-step PCR. Step 1 (Amplification): Use 5-10µg of gDNA per sample in a 100µL reaction with primers containing partial Illumina adapters. Run 18-22 cycles to amplify the sgRNA cassette. Step 2 (Indexing): Use 1µL of purified PCR1 product in a 50µL reaction with unique dual indexing primers for each sample (8-10 cycles).
- Critical: Split PCR1 reactions across multiple tubes to avoid amplification bias from excessive cycles.
- Pooling & Sequencing: Quantify all PCR2 products by qPCR (KAPA Library Quant Kit) and pool equimolarly. Sequence on an Illumina NextSeq 550/2000 using a 75bp single-end run. Aim for 200-500 reads per original cell.
Visualization of Workflow & Statistical Concepts
Title: Powered CRISPRa/i Screen Experimental Workflow
Title: Key Factors Influencing Statistical Power
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagent Solutions for Powered CRISPRa/i Screens
| Reagent / Material | Function & Critical Specification | Example Product/Catalog |
|---|---|---|
| Focused CRISPRa/i Library | Pooled sgRNA library targeting gene sets (e.g., druggable genome, transcription factors) with 6-10 sgRNAs/gene and high on-target scores. Must include non-targeting control guides. | Calabrese et al. (Nature Biotech, 2023) human CRISPRa-v2 library; Dolcetto et al. (Cell, 2022) genome-wide CRISPRi library. |
| Lentiviral Packaging Mix | For high-titer, replication-incompetent lentivirus production. 3rd generation systems (psPAX2, pMD2.G) are standard. Must ensure low recombination risk. | MERCK Sigma Lenti-Vpak, Invitrogen ViraPower Lentiviral Packaging Mix. |
| Large-Scale gDNA Extraction Kit | For high-yield, high-quality genomic DNA from 10^7 to 10^8 cells. Purity (A260/280) is critical for PCR efficiency. | Qiagen Blood & Cell Culture DNA Maxi Kit (Cat. #13362), NucleoBond HEF Maxi. |
| High-Fidelity PCR Master Mix | For unbiased, low-error amplification of sgRNA sequences from gDNA during NGS library prep. Minimal amplification bias is essential. | NEB Next Ultra II Q5 Master Mix, KAPA HiFi HotStart ReadyMix. |
| Dual-Indexed Sequencing Primers | Unique combinatorial indexes for multiplexing many samples (T0, T-end, multiple replicates) in a single sequencing run, enabling accurate demultiplexing. | Illumina Nextera XT Index Kit v2, IDT for Illumina UD Indexes. |
| Cell Selection Antibiotic | To select for stable sgRNA expression. Puromycin is most common. Must titrate to determine minimal effective concentration for 100% kill in 3-5 days on non-transduced cells. | Thermo Fisher Scientific Puromycin Dihydrochloride. |
| Library Quantification Kit | Accurate quantification of the final pooled NGS library by qPCR for precise loading onto the sequencer. | KAPA Library Quantification Kit for Illumina platforms. |
Within CRISPR activation (CRISPRa) and interference (CRISPRi) screening for gene function research, robust validation is paramount. These screens systematically overexpress or repress genes to identify hits involved in phenotypes like drug resistance or cell differentiation. However, significant noise and false positives necessitate stringent controls and benchmarks. This article details essential validation strategies, protocols, and reagents to ensure screen reliability and reproducibility for research and drug development.
Core Validation Controls: Types and Implementation
Effective screen validation employs multiple control types to address specific confounding factors.
Table 1: Essential Control Types for CRISPRa/i Screens
| Control Type | Purpose in CRISPRa/i | Typical Implementation | Data Output |
|---|---|---|---|
| Non-Targeting Controls (NTCs) | Define baseline signal, assess off-target effects. | sgRNAs with no known genomic target. | Distribution of negative phenotype scores. |
| Essential Gene Positive Controls | Confirm screen efficacy for repression (CRISPRi). | sgRNAs targeting core essential genes (e.g., ribosomal proteins). | Significant depletion in survival screens. |
| Non-Essential Gene Negative Controls | Confirm screen specificity for repression (CRISPRi). | sgRNAs targeting "safe harbor" or non-essential loci. | No depletion in survival screens. |
| Expression Controls | Confirm activation (CRISPRa) or repression (CRISPRi) mechanism. | sgRNAs targeting a constitutive reporter (e.g., GFP). | Flow cytometry measurement of reporter signal. |
| Pool Balancing Controls | Monitor sgRNA representation and library quality. | Use of non-functional or scrambled sgRNAs in constant abundance. | Sequencing count stability over passages. |
Benchmarking with Reference Gene Sets
Quantitative benchmarking against established gene sets transforms qualitative assessment into a metric-driven process.
Table 2: Common Benchmarking Gene Sets for Functional Screens
| Benchmark Set | Application | Expected Phenotype (CRISPRi) | Validation Metric |
|---|---|---|---|
| Core Essential Genes (e.g., Hart et al.) | Assay sensitivity for dropout screens. | Strong depletion. | Recall (e.g., AUC of ROC curve). |
| Common Essential Genes (DepMap) | Assay sensitivity in specific cell line. | Strong depletion. | Gene-level ROC AUC or precision-recall. |
| Non-Essential Genes (DepMap) | Assay specificity, false positive rate. | No depletion. | False Discovery Rate (FDR) at given threshold. |
| Pathway-Specific Genes (e.g., Proteasome) | Assay biological relevance. | Coordinated dropout or enrichment. | Gene Set Enrichment Analysis (GSEA) score. |
Detailed Experimental Protocols
Protocol 1: Validation of Screen Performance Using Essential/Non-Essential Benchmarks
Objective: Quantify the sensitivity and specificity of a CRISPRi dropout screen. Materials: Infected cell pool post-selection, genomic DNA extraction kit, PCR reagents, NGS sequencing platform. Procedure:
- Cell Culture & Harvest: Culture the transduced cell pool for 14 population doublings. Harvest 1e7 cells at passages corresponding to ~5 and ~14 doublings for gDNA extraction.
- gDNA Extraction & Amplification: Extract gDNA using a large-scale kit. Amplify sgRNA libraries via two-step PCR (Step 1: amplify integrated sgRNA; Step 2: add Illumina adapters and sample indexes). Use at least 500ng gDNA per PCR reaction to maintain library diversity.
- Sequencing & Read Alignment: Sequence on an Illumina NextSeq (75bp single-end). Align reads to the reference sgRNA library using Bowtie2 or a similar aligner. Count reads per sgRNA.
- Data Analysis: Normalize counts using median scaling. Calculate log2(fold-change) between passage 14 and passage 5 for each sgRNA. Aggregate sgRNA scores to gene scores (e.g., using MAGeCK or CERES).
- Benchmarking: Compare gene scores to reference essential/non-essential lists. Calculate the Area Under the Curve (AUC) of the Receiver Operating Characteristic (ROC) curve. A high AUC (>0.8) indicates strong screen performance.
Protocol 2: Validation of CRISPRa/i Modulation via Flow Cytometry
Objective: Confirm transcriptional activation or repression at a single-cell level. Materials: Reporter cell line with integrated GFP under a minimal promoter, lentiviral delivery vectors for CRISPRa/i sgRNAs, flow cytometer. Procedure:
- Reporter Line Engineering: Generate a stable cell line with a TRE-minimal promoter-GFP construct. For CRISPRa, deliver an sgRNA targeting the TRE promoter. For CRISPRi, deliver an sgRNA targeting the GFP open reading frame.
- Transduction & Selection: Transduce reporter cells at low MOI (<0.3) and select with appropriate antibiotics for 5-7 days.
- Analysis: Harvest cells, resuspend in PBS + 2% FBS, and analyze on a flow cytometer. Use cells transduced with a non-targeting sgRNA (NTC) as the baseline. A successful CRISPRa experiment should show a right-shift in GFP fluorescence; CRISPRi should show a left-shift.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for CRISPRa/i Screen Validation
| Item | Function & Application | Example Product/Kit |
|---|---|---|
| Validated sgRNA Library | Pre-designed library with targeting and control sgRNAs. | Toronto KnockOut (TKO) v3 for essentiality; Calabrese et al. CRISPRa/i libraries. |
| gDNA Extraction Kit | High-yield, purification of genomic DNA from large cell pellets. | Qiagen Blood & Cell Culture DNA Maxi Kit. |
| PCR Master Mix | High-fidelity amplification of sgRNA sequences from gDNA. | KAPA HiFi HotStart ReadyMix. |
| NGS Indexing Primers | Addition of unique dual indexes for sample multiplexing. | Illumina TruSeq CD Indexes. |
| Flow Cytometry Antibodies | Detection of surface markers for secondary validation. | BioLegend Anti-CD44, Anti-CD133. |
| Cell Viability Assay | Quantification of dropout/enrichment phenotypes. | CellTiter-Glo Luminescent Cell Viability Assay. |
Visualization of Workflows and Pathways
Title: CRISPR Screen Validation and Benchmarking Workflow
Title: CRISPRa and CRISPRi Mechanism Diagram
Validating Hits and Comparing Technologies: From Screen Data to Confident Discovery
Within the broader thesis on utilizing CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screens for gene activation and repression research, robust bioinformatic analysis is critical for hit identification. This article provides detailed application notes and protocols for three essential computational tools: MAGeCK for screen analysis, PinAPL-Py for pooled screen analysis, and CRISPResso2 for editing efficiency quantification. These pipelines enable researchers to transition from raw sequencing data to high-confidence gene targets, crucial for functional genomics and early-stage drug discovery.
Application Notes & Protocols
MAGeCK (Model-based Analysis of Genome-wide CRISPR-Cas9 Knockout)
Application Note: MAGeCK is a comprehensive computational tool designed for the analysis of both positive and negative selection screens. It is particularly valuable in CRISPRi/a screens for identifying genes whose repression or activation significantly impacts a phenotype of interest (e.g., cell viability, fluorescence intensity). It robustly handles variability across replicates and computes significance using a negative binomial model.
Key Protocol: Analysis of a CRISPRi Screen for Essential Genes
- Data Input Preparation: Prepare a raw count table (sgRNA per sample) and a sample annotation file linking samples to conditions (e.g., T0, T14). Prepare the library file specifying sgRNA-gene mappings.
- Quality Control: Run
mageck countto generate raw counts from FASTQ files and perform initial QC. - Test for Positive/Negative Selection: Run
mageck testto compare conditions (e.g., post-selection vs. initial plasmid). - Visualization & Hit Calling: Generate rank plots and visualize top hits. Genes with a negative beta score and FDR < 0.05 are candidate essential genes (for CRISPRi).
Table 1: Representative MAGeCK Output for a CRISPRi Viability Screen
| Gene | sgRNA # | Beta Score | p-value | FDR | Interpretation |
|---|---|---|---|---|---|
| POLR2D | 5 | -3.21 | 2.5E-08 | 1.1E-05 | Core essential gene |
| MYC | 4 | -1.85 | 7.3E-04 | 0.012 | Context-dependent essential |
| CDK4 | 5 | -0.32 | 0.45 | 0.67 | Not significant |
PinAPL-Py (Pooled in-vitro and in-vivo Analysis Platform - Python)
Application Note: PinAPL-Py is a flexible platform for analyzing complex pooled screen data, including time-course and in-vivo experiments. It excels in normalizing read counts, calculating fold-changes, and generating gene-level scores. Its strength in longitudinal data analysis makes it suitable for tracking gene effects over time in CRISPRa/i screens.
Key Protocol: Time-Course Analysis of a CRISPRa Screen
- Data Normalization: Input raw sgRNA count files for each time point. Normalize counts to reads per million (RPM) and perform log2 transformation.
- Fold-Change Calculation: Compute fold-changes for each sgRNA relative to the T0 reference sample for each subsequent time point.
- Gene Score Calculation: Use the PIN tool (PINAPL-Py's core) to aggregate sgRNA fold-changes into a robust gene-level score (e.g., using a weighted average).
- Hit Identification: Rank genes by their cumulative or time-point-specific scores. Apply thresholds (e.g., top/bottom 5% of distribution) to identify hits.
Table 2: PinAPL-Py Output for a 3-Week CRISPRa Time-Course Screen
| Gene | Week 1 Score | Week 2 Score | Week 3 Score | Trend | Final Rank |
|---|---|---|---|---|---|
| IL6R | 0.55 | 1.32 | 2.87 | Increasing | 1 |
| EGFR | 1.21 | 1.45 | 1.50 | Stable High | 5 |
| TP53 | -0.02 | -0.11 | -0.15 | Stable Neutral | 4500 |
CRISPResso2
Application Note: CRISPResso2 quantifies genome editing efficiency from next-generation sequencing data. In CRISPRa/i screens, it is vital for QC of the screening library, confirming the intended epigenetic perturbation (via verification of gRNA binding site integrity in the absence of indels) and validating hits in follow-up experiments.
Key Protocol: Validation of CRISPRi sgRNA Activity
- Amplicon Sequencing: Design primers to amplify the genomic region targeted by the sgRNA from validation cell pools. Perform NGS.
- Alignment and Quantification: Run CRISPResso2 to align reads to the reference amplicon and quantify indels and modifications.
- Interpretation: For a valid CRISPRi hit, expect high read alignment (>80%) but very low indel frequency (<5%), confirming repression is due to dCas9 binding/recruitment rather than sequence disruption.
Table 3: CRISPResso2 Output for Hit Validation
| Sample | Total Reads | Aligned % | % Unmodified | % Indels | % Other Modifications |
|---|---|---|---|---|---|
| Non-Targeting Ctrl | 150,000 | 98.2 | 97.5 | 1.2 | 1.3 |
| Candidate Hit A | 145,500 | 85.4 | 84.1 | 1.0 | 14.9* |
| Candidate Hit B | 138,750 | 97.8 | 48.2 | 46.5 | 5.3 |
*Likely represents heterozygous SNP or sequencing error; confirms low editing, suitable for CRISPRi.
The Scientist's Toolkit
Table 4: Key Research Reagent Solutions for CRISPRa/i Screen Analysis
| Item | Function in Analysis Pipeline |
|---|---|
| sgRNA Library Plasmid Pool | Physical source of sgRNA sequences; defines screen targets and controls. |
| High-Quality NGS Library Prep Kit | Prepares sequencing libraries from PCR-amplified sgRNA inserts or genomic amplicons. |
| Bowtie2 or BWA Aligners | Maps sequencing reads to the reference sgRNA library or genome. |
| MAGeCK RRA Algorithm | Ranks genes based on sgRNA enrichment/depletion patterns robustly. |
| PinAPL-Py PIN Tool | Aggregates sgRNA-level fold-changes into gene scores across complex experiments. |
| CRISPResso2 Allele Frequency Table | Quantifies the spectrum and frequency of editing events at target loci. |
| Negative Control sgRNAs (e.g., Targeting GFP) | Provide a baseline for calculating fold-changes and statistical significance. |
| Positive Control sgRNAs (e.g., Targeting Essential Genes) | Assess screen dynamic range and performance in selection assays. |
Visualizations
Title: MAGeCK Analysis Workflow for CRISPR Screens
Title: CRISPResso2 Editing Analysis Pipeline
Title: Pipeline Integration in CRISPRa/i Thesis Research
Following a pooled or arrayed CRISPR activation (CRISPRa) or interference (CRISPRi) screen for gene function, candidate hits require rigorous validation. This involves a tiered strategy: primary validation (RT-qPCR for mRNA, Western Blot for protein) confirms the direct molecular outcome of the perturbation. Secondary validation (functional assays) confirms the expected phenotypic consequence, establishing a direct link between the genetic perturbation, target expression, and cellular phenotype.
Primary Validation: Molecular Confirmation
RT-qPCR Protocol for mRNA Level Validation
Aim: Quantify changes in target gene mRNA expression following CRISPRa or CRISPRi perturbation.
Detailed Protocol:
- Cell Harvest & Lysis: 72 hours post-transduction/transfection, harvest ~1x10^6 cells. Lyse using a monophasic reagent (e.g., TRIzol).
- RNA Isolation: Perform phase separation with chloroform. Precipitate RNA with isopropanol, wash with 75% ethanol, and resuspend in RNase-free water.
- DNase Treatment: Treat 1 µg of total RNA with DNase I to remove genomic DNA contamination.
- cDNA Synthesis: Use a high-capacity cDNA reverse transcription kit with random hexamers.
- qPCR Setup: Prepare reactions in triplicate using a SYBR Green or TaqMan master mix.
- Primer Design: Design primers spanning an exon-exon junction. Validate efficiency (90-110%).
- Reaction: 10 µL SYBR Green mix, 0.5 µM each primer, 2 µL cDNA (diluted 1:10), nuclease-free water to 20 µL.
- qPCR Run: Use standard cycling conditions (95°C for 3 min, then 40 cycles of 95°C for 10 sec, 60°C for 30 sec).
- Data Analysis: Calculate ∆∆Ct values using at least two validated reference genes (e.g., GAPDH, ACTB). Report fold-change relative to non-targeting sgRNA control.
Table 1: Example RT-qPCR Data from a CRISPRa Screen Hit Validation
| Target Gene | Condition (sgRNA) | Mean ∆Ct (vs. Ref) | Fold-Change (vs. NT) | p-value (t-test) |
|---|---|---|---|---|
| Gene X | NT Control | 5.2 | 1.0 | - |
| CRISPRa_1 | 3.1 | 4.2 | 0.003 | |
| CRISPRa_2 | 3.4 | 3.5 | 0.008 | |
| Gene Y | NT Control | 7.8 | 1.0 | - |
| CRISPRi_1 | 9.5 | 0.3 | 0.001 | |
| CRISPRi_2 | 10.1 | 0.2 | 0.0005 |
NT: Non-Targeting; Ref: Reference Genes (GAPDH, ACTB).
Western Blot Protocol for Protein Level Validation
Aim: Confirm changes in target protein abundance or post-translational modification.
Detailed Protocol:
- Cell Lysis: 96-120 hours post-perturbation, lyse cells in RIPA buffer + protease/phosphatase inhibitors on ice for 30 min. Centrifuge (14,000g, 15 min, 4°C).
- Protein Quantification: Determine supernatant concentration using a BCA assay.
- Sample Preparation: Mix 20-30 µg protein with Laemmli buffer, denature at 95°C for 5 min.
- Gel Electrophoresis: Load samples onto a 4-20% gradient SDS-PAGE gel. Run at 120V for ~90 min.
- Transfer: Transfer proteins to a PVDF membrane using a wet or semi-dry system.
- Blocking: Block membrane with 5% non-fat milk in TBST for 1 hour at RT.
- Antibody Incubation:
- Primary Antibody: Incubate with target-specific antibody (e.g., 1:1000 dilution in 5% BSA/TBST) overnight at 4°C.
- Wash: 3 x 10 min with TBST.
- Secondary Antibody: Incubate with HRP-conjugated antibody (1:5000 in blocking buffer) for 1 hour at RT. Wash again.
- Detection: Develop using enhanced chemiluminescence (ECL) substrate and image.
- Stripping & Re-probing: Strip membrane (optional) and re-probe for a loading control (e.g., GAPDH, β-Actin, Vinculin).
Table 2: Key Reagents for Primary Validation
| Reagent / Solution | Function in Protocol | Critical Parameters / Notes |
|---|---|---|
| dCas9-VP64/SAM (CRISPRa) or dCas9-KRAB (CRISPRi) System | Engineered CRISPR complex for transcriptional activation or repression. | Ensure optimal sgRNA design for the intended system (e.g., near TSS for CRISPRi). |
| TRIzol / Monophasic Lysis Reagent | Simultaneous lysis and stabilization of RNA, DNA, and protein. | For RT-qPCR, maintain RNase-free conditions during isolation. |
| SYBR Green or TaqMan qPCR Master Mix | Contains enzymes, dNTPs, and dye for real-time PCR quantification. | SYBR Green requires specific primers; TaqMan uses a probe for higher specificity. |
| Validated qPCR Primers | Amplify specific cDNA target sequence. | Must be exon-spanning, with 90-110% efficiency. Test for specificity via melt curve. |
| RIPA Lysis Buffer | Cell lysis and protein extraction for Western Blot. | Must be supplemented with fresh protease/phosphatase inhibitors. |
| Target-Specific Primary Antibody | Binds specifically to the protein of interest. | Validate for application (Western Blot) and species reactivity. Check literature for citations. |
| HRP-conjugated Secondary Antibody | Binds primary antibody for chemiluminescent detection. | Must be raised against the host species of the primary antibody (e.g., anti-rabbit). |
Secondary Validation: Functional Phenotypic Confirmation
Aim: To link the molecular change (validated by RT-qPCR/WB) to a relevant phenotypic output from the original screen (e.g., proliferation, migration, reporter activity).
Functional Assay Protocol: Luciferase Reporter Assay
Aim: Validate transcriptional regulation of a specific pathway or promoter element.
Protocol:
- Reporter Construct: Clone the putative promoter or response element of a downstream target gene into a luciferase reporter vector (e.g., pGL4).
- Co-transfection: Co-transfect cells with (a) the dCas9-effector (CRISPRa/i), (b) the validated sgRNA, and (c) the luciferase reporter construct. Include a Renilla luciferase control plasmid for normalization.
- Incubation: Culture cells for 48-72 hours.
- Lysis & Measurement: Lyse cells with Passive Lysis Buffer. Measure firefly and Renilla luciferase activity sequentially using a dual-luciferase assay kit on a plate reader.
- Analysis: Normalize firefly luminescence to Renilla luminescence for each well. Compare relative light units (RLU) between targeting and non-targeting sgRNA conditions.
Table 3: Example Functional Validation Data
| Assay Type | Target Gene | Perturbation | Measured Output (vs. NT Control) | p-value | Links to Screen Phenotype? |
|---|---|---|---|---|---|
| Luciferase Reporter | Gene A (a TF) | CRISPRa | 5.8x RLU Increase | 0.002 | Validated increased pathway activity from original screen. |
| Proliferation (Cell Titer-Glo) | Oncogene B | CRISPRi | 45% Viability Decrease | <0.0001 | Confirms screen hit essential for growth. |
| Transwell Migration | Metastasis Gene C | CRISPRi | 70% Reduction in Migrated Cells | 0.005 | Validates role in invasion/migration phenotype. |
Visualizations
Tiered Validation Workflow for CRISPR Screens
RT-qPCR Workflow for mRNA Validation
Western Blot Workflow for Protein Validation
Functional genomics screens are pivotal for identifying genes involved in biological processes and therapeutic target discovery. This application note compares four principal technologies used for perturbation screens: CRISPR activation/interference (CRISPRa/i), RNA interference (RNAi), CRISPR knockout (CRISPR-KO), and Small Molecule Screens. Framed within the context of a broader thesis on CRISPRa/i screens for gene activation/repression research, we detail their mechanisms, applications, and provide protocols for implementation.
Technology Comparison: Mechanisms & Applications
Core Mechanism of Action
- CRISPRa (Activation): Uses a catalytically dead Cas9 (dCas9) fused to transcriptional activation domains (e.g., VP64, p65, Rta) to recruit RNA polymerase II and co-activators to specific gene promoters, upregulating transcription.
- CRISPRi (Interference): Uses dCas9 fused to transcriptional repressor domains (e.g., KRAB, SID4x) to block RNA polymerase binding or elongation, downregulating transcription.
- RNAi: Utilizes short-hairpin RNAs (shRNAs) or siRNAs that are loaded into the RNA-induced silencing complex (RISC) to guide cleavage and degradation of complementary mRNA sequences.
- CRISPR-KO: Employs Cas9 nuclease to create targeted double-strand breaks (DSBs), leading to frameshift mutations via non-homologous end joining (NHEJ) and permanent gene knockout.
- Small Molecule Screens: Use libraries of chemical compounds that modulate protein function through inhibition, activation, or stabilization, offering temporal control and dose-responsiveness.
Quantitative Performance Comparison
Table 1: Comparative Overview of Screening Technologies
| Feature | CRISPRa | CRISPRi | CRISPR-KO | RNAi | Small Molecules |
|---|---|---|---|---|---|
| Primary Use | Gain-of-function | Loss-of-function (Transcriptional) | Loss-of-function (Genetic) | Loss-of-function (Transcriptional) | Pharmacological Modulation |
| Target | DNA (Promoter) | DNA (Promoter/TSS) | DNA (Coding Exon) | mRNA | Protein |
| Effect | Transcriptional Activation | Transcriptional Repression | Permanent Gene Disruption | mRNA Degradation/Translational Block | Protein Inhibition/Activation |
| On-Target Efficiency | High (>80% induction common) | High (>70% repression common) | Very High (near-complete KO) | Variable (often 70-90% knockdown) | Highly Compound-Dependent |
| Off-Target Effects | Low (high DNA specificity) | Low (high DNA specificity) | Moderate (DNA cleavage specificity) | High (seed-based miRNA-like effects) | Variable (binding promiscuity) |
| Temporal Control | Moderate (depends on delivery) | Moderate (depends on delivery) | None (Permanent) | Moderate (transient siRNA) | Excellent (dose/time) |
| Screen Readiness | Excellent (lentiviral libraries) | Excellent (lentiviral libraries) | Excellent (Gold Standard) | Good (lentiviral libraries) | Excellent (commercial libraries) |
| Therapeutic Link | Direct (gene therapy) | Direct (gene therapy) | Indirect (target ID) | Indirect (target ID) | Direct (drug discovery) |
Table 2: Typical Screen Metrics (Pooled Library Format)
| Parameter | CRISPRa/i | CRISPR-KO | RNAi |
|---|---|---|---|
| Library Size (Human) | ~50,000 - 200,000 sgRNAs | ~50,000 - 120,000 sgRNAs | ~50,000 - 150,000 shRNAs |
| sg/shRNAs per Gene | 3-10 (multiple promoter targets) | 3-10 (multiple exonic targets) | 3-10 (multiple mRNA targets) |
| Screen Duration | 2-4 weeks (plus assay) | 2-4 weeks (plus assay) | 2-3 weeks (plus assay) |
| Key Validation Step | RT-qPCR / Western Blot | Sanger Sequencing / T7E1 | RT-qPCR / Western Blot |
Detailed Experimental Protocols
Protocol: Pooled CRISPRa/i Screen for Resistance/Sensitivity
Objective: Identify genes whose transcriptional activation (CRISPRa) or repression (CRISPRi) confer resistance or sensitivity to a drug treatment.
Part A: Library Lentivirus Production
- Plate HEK293T cells in 15cm dishes (70-80% confluent) in DMEM + 10% FBS, 1-2 hours before transfection.
- Prepare Transfection Mix (per dish):
- 22.5 µg CRISPRa/i sgRNA library plasmid (e.g., Calabrese, SAM, or CRISPRi-v2)
- 16.5 µg psPAX2 packaging plasmid
- 6 µg pMD2.G envelope plasmid
- 135 µL 1M CaCl2
- Add nuclease-free water to 1.125 mL total.
- Add 1.125 mL of 2X HEPES-Buffered Saline (pH 7.05) dropwise while vortexing.
- Incubate 5 minutes at room temperature.
- Add mixture dropwise to HEK293T cells. Replace medium after 6-8 hours.
- Harvest virus at 48 and 72 hours post-transfection. Pool supernatants, filter through 0.45µm PES filter, and concentrate via ultracentrifugation (50,000 x g, 2h, 4°C) or using Lenti-X concentrator. Aliquot and store at -80°C. Titer virus on target cells.
Part B: Cell Infection and Screening
- Determine MOI: Perform a pilot infection with a non-targeting sgRNA virus. Aim for MOI ~0.3-0.4 to ensure most cells receive a single sgRNA.
- Library Infection: Scale up to infect >500 cells per sgRNA in the library to maintain representation. For a 50,000 sgRNA library, infect at least 2.5x10^7 cells. Add polybrene (8µg/mL final).
- Selection: Begin puromycin selection (1-3µg/mL, dependent on cell line) 48 hours post-infection. Maintain selection for 5-7 days.
- Perturbation: Split cells into treatment (drug) and vehicle control arms. Maintain cells for 14-21 population doublings, ensuring >500x coverage per sgRNA is maintained at each passage.
- Harvest Genomic DNA: Pellet at least 1x10^7 cells per arm at endpoint. Use a commercial gDNA extraction kit (e.g., Qiagen Blood & Cell Culture DNA Maxi Kit). Elute in TE buffer.
Part C: NGS Library Prep & Analysis
- PCR Amplify sgRNA inserts:
- Round 1 (18 cycles): Amplify 5-10 µg gDNA per sample in 100µL reactions using Herculase II polymerase and primers adding partial Illumina adapter sequences.
- Purify PCR products (AMPure XP beads).
- Round 2 (12 cycles): Index PCR to add full Illumina adapters and sample barcodes.
- Purify final library, quantify by qPCR, and pool for sequencing. Aim for >100 reads per sgRNA.
- Sequencing & Analysis: Sequence on an Illumina NextSeq (75bp single-end). Align reads to the sgRNA library reference file. Use MAGeCK or PinAPL-Py algorithms to compare sgRNA abundances between treatment and control arms, identifying significantly enriched or depleted sgRNAs (FDR < 0.1).
Protocol: CRISPR-KO Screen (Counter-Screen Validation)
Objective: Validate hits from a CRISPRa/i screen by performing a loss-of-function counter-screen using CRISPR-KO.
- Design Validation Library: Select top 50-100 gene hits. Design 5 sgRNAs per gene targeting early coding exons using tools like CRISPick or CHOPCHOP. Include 20 non-targeting controls.
- Clone into LentiGuide-Puro: Perform arrayed cloning or pooled oligo synthesis/cloning.
- Execute Screen: Follow the pooled screen protocol in 3.1, Part B & C, but use the validation KO library. Expected Result: Genes whose activation conferred drug resistance should show the opposite phenotype (sensitivity) when knocked out, confirming the specificity of the original hit.
Protocol: RNAi Screen (Comparative Orthology)
Objective: Compare results from a CRISPRi screen with an RNAi screen for transcriptional repression.
- Library: Use a commercially available genome-wide shRNA library (e.g., TRC, Decipher).
- Virus Production & Infection: Similar to Protocol 3.1A/B, but using shRNA library plasmids and appropriate packaging systems.
- Key Difference: Screen duration is typically shorter (10-14 days) due to transient nature of knockdown. Harvest cells for genomic DNA (to amplify the shRNA barcode) or directly extract RNA for phenotypic readouts.
- Analysis: Sequence shRNA barcodes and analyze similarly. Note increased off-target effects; require multiple shRNAs per gene for confidence.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for CRISPRa/i and Comparative Screens
| Reagent / Material | Function & Description | Example Product (Vendor) |
|---|---|---|
| dCas9-VPR / dCas9-KRAB Expression Plasmid | Stable expression of the CRISPRa or CRISPRi effector protein. | pHAGE EF1α dCas9-VPR (Addgene #114257); lenti dCas9-KRAB-blast (Addgene #89567) |
| sgRNA Library Lentiviral Plasmid | Pooled vector encoding sgRNAs targeting the genome. | Calabrese CRISPRa Lib (Addgene #125054); Human CRISPRi-v2 (Addgene #83979) |
| Lentiviral Packaging Plasmids | Required for production of replication-incompetent lentivirus. | psPAX2 (Addgene #12260); pMD2.G (Addgene #12259) |
| HEK293T/293FT Cell Line | Highly transfectable cell line for high-titer lentivirus production. | HEK293T/17 (ATCC CRL-11268) |
| Polybrene (Hexadimethrine bromide) | A cationic polymer that enhances viral transduction efficiency. | Sigma-Aldrich H9268 |
| Puromycin Dihydrochloride | Selective antibiotic for cells expressing a puromycin resistance gene (PuroR). | Thermo Fisher Scientific A1113803 |
| Genomic DNA Extraction Kit | For high-yield, high-quality gDNA from large cell pellets for sgRNA recovery. | QIAGEN Blood & Cell Culture DNA Maxi Kit (13362) |
| Herculase II Fusion DNA Polymerase | High-fidelity polymerase for robust amplification of sgRNA sequences from gDNA. | Agilent 600679 |
| SPRIselect Beads (AMPure XP) | For size-selective purification and cleanup of PCR-amplified NGS libraries. | Beckman Coulter B23319 |
| Next-Generation Sequencer | Platform for deep sequencing of sgRNA amplicons. | Illumina NextSeq 550/2000 |
| Screen Analysis Software | Computational tool for identifying significantly enriched/depleted sgRNAs. | MAGeCK (https://sourceforge.net/p/mageck) |
Data Interpretation & Strategic Selection Guide
Table 4: Technology Selection Matrix Based on Research Goal
| Research Goal | Recommended Primary Technology | Key Rationale | Recommended Counter-Screen |
|---|---|---|---|
| Identify Genes Conferring Drug Resistance | CRISPRa | Unbiased discovery of activation-driven resistance mechanisms. | CRISPR-KO or CRISPRi |
| Identify Essential Genes (Lethality) | CRISPR-KO | Highest on-target efficiency for complete loss-of-function. | CRISPRi (for hypomorph) |
| Identify Synthetic Lethal Interactions | CRISPRi | Tunable, partial knockdown better models therapeutic inhibition. | CRISPR-KO (for validation) |
| Rapid, Acute Protein Inhibition | Small Molecules | Excellent temporal control and dose response. | CRISPRi/KO (for genetic validation) |
| High-Throughput Target Discovery | Pooled CRISPR-KO | Gold standard for genome-wide loss-of-function screens. | RNAi (orthogonal validation) |
| Studies with Potential Off-Target Concerns | CRISPRa/i | Highest DNA-specificity; minimal off-target transcriptional effects. | Use multiple sgRNAs per gene |
Conclusion: The choice between CRISPRa/i, CRISPR-KO, RNAi, and small molecule screens is dictated by the biological question, desired perturbation type (activation, complete KO, knockdown, or pharmacological), and required specificity. CRISPRa/i screens offer precise transcriptional modulation with high specificity, filling a critical niche between permanent KO and transient small-molecule effects, making them indispensable for comprehensive functional genomics research and drug target discovery pipelines.
Integrating CRISPRa and CRISPRi Data for Complementary Gene Function Insights
Within the broader thesis on CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) screens for gene activation and repression research, the integration of these complementary modalities is a powerful strategy. By combining loss-of-function (CRISPRi) and gain-of-function (CRISPRa) data from parallel screens, researchers can obtain a more complete picture of gene function, identify context-specific dependencies, and mitigate false positives. This application note provides protocols and frameworks for the design, execution, and integrated analysis of dual CRISPRa/i screens.
Key Advantages of Integrated Analysis
The synergistic use of CRISPRa and CRISPRi data offers several critical insights:
- Validation of Hit Specificity: Genes whose perturbation (activation or repression) consistently produces a phenotype are high-confidence hits.
- Directional Phenotype Elucidation: Opposite phenotypic directions from CRISPRa vs. CRISPRi for the same gene confirm the specificity of the observed effect (e.g., activation rescues a defect while repression exacerbates it).
- Identification of Buffering Genes: Genes where only one modality (a or i) produces a phenotype may reveal buffered pathways or synthetic lethal interactions.
- Mapping of Gene Regulatory Networks: Integrated data can inform models of epistatic relationships and pathway architecture.
Application Notes: Design and Analysis
Experimental Design Considerations
- Library Design: Use genome-wide or focused libraries with matched sgRNAs for both CRISPRa and CRISPRi targeting the same gene sets. Ensure transcriptional start site (TSS) targeting for CRISPRi and promoter-proximal targeting for CRISPRa.
- Cell Models: Utilize cell lines expressing both a reverse tetracycline-controlled transactivator (rtTA) for inducible dCas9-effector expression (e.g., dCas9-KRAB for i; dCas9-VPR for a) and a constitutively active Cas9 for knockout controls.
- Screen Conditions: Perform parallel screens under identical selective pressures (e.g., drug treatment, nutrient deprivation, viral infection). Include appropriate controls (non-targeting sgRNAs, essential/positive control genes).
Integrated Data Analysis Workflow
A generalized bioinformatics pipeline for integrated analysis is depicted below.
Title: Integrated CRISPRa/i Data Analysis Workflow
Quantitative Data Integration and Interpretation
Summarize gene-level scores from both screens into a comparative table. Key metrics include log2 fold change (LFC), p-value, false discovery rate (FDR), and a combined confidence score.
Table 1: Exemplar Integrated Hits from a Dual CRISPRa/i Survival Screen
| Gene | CRISPRi LFC | CRISPRi FDR | CRISPRa LFC | CRISPRa FDR | Phenotype Concordance* | Integrated Confidence |
|---|---|---|---|---|---|---|
| MYC | -2.3 | 1.2E-10 | +1.8 | 3.5E-08 | Opposite (Validated) | High |
| KRAS | -1.9 | 5.7E-09 | +0.4 | 0.32 | i-Only (Essential) | High |
| TP53 | -0.6 | 0.07 | +2.1 | 1.1E-06 | a-Only (Suppressor) | Medium |
| BRD4 | -3.1 | 2.4E-12 | -1.2 | 0.04 | Concordant (Complex) | Requires Validation |
| ACTB | +0.1 | 0.65 | -0.1 | 0.71 | None (Negative Ctrl) | Low |
Phenotype Concordance: "Opposite" = a and i show significant, opposing phenotypes; "i-Only/a-Only" = only one modality is significant; "Concordant" = both a and i show phenotype in same direction. *Integrated Confidence: Heuristic based on significance, effect size, and concordance pattern.
Detailed Protocols
Protocol 1: Running Paired CRISPRa and CRISPRi Screens
Objective: To identify genes affecting resistance to a targeted oncology compound (e.g., a BRAF inhibitor).
Materials: See "The Scientist's Toolkit" below.
Method:
- Cell Line Preparation: Generate a polyclonal population of the desired cell line (e.g., A375 melanoma) stably expressing dCas9-KRAB (for i) or dCas9-VPR (for a) under a doxycycline-inducible promoter. Validate effector expression and inducibility via immunoblot.
- Viral Transduction: For each screen arm (a and i), independently transduce cells with the respective lentiviral sgRNA library (e.g., Calabrese whole-genome library) at a low MOI (<0.3) to ensure single integration. Use 500x library coverage.
- Selection and Induction: Puromycin-select transduced cells for 5-7 days. Induce dCas9-effector expression with doxycycline (1 µg/mL) 48 hours prior to selection pressure.
- Screen Execution: Split cells into treated (BRAF inhibitor) and untreated (DMSO) control arms. Maintain cells for 14-21 population doublings, ensuring minimum 500x coverage is maintained throughout.
- Harvest and Sequencing: Harvest genomic DNA from final populations and the initial plasmid library. Amplify sgRNA sequences via PCR and subject to next-generation sequencing (NGS) on an Illumina platform.
Protocol 2: Integrated Computational Analysis
Objective: To analyze sequencing data from Protocol 1 and generate an integrated hit list.
Software: MAGeCK (version 0.5.9+), R/Bioconductor, Python (Pandas, NumPy).
Method:
- Read Processing: Demultiplex NGS reads and align to the sgRNA library reference file using
mageck count.
Separate Screen Analysis: Run MAGeCK MLE separately for CRISPRi and CRISPRa data to calculate gene-level β scores (LFC) and FDRs.
Data Integration: In R, merge the two result tables by gene symbol. Calculate a combined score (e.g., product of signed -log10(FDR) from each screen). Filter for genes where
FDR < 0.1in at least one screen.- Directional Phenotype Plot: Create a scatter plot of CRISPRi LFC vs. CRISPRa LFC. Color-code points based on significance and concordance patterns (see Diagram 2).
- Pathway Enrichment: Perform Gene Set Enrichment Analysis (GSEA) on the combined score ranking using the
fgseapackage to identify affected pathways.
Title: Interpretation of CRISPRa/i Integration Scatter Plot
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function | Example Product/Catalog |
|---|---|---|
| dCas9-Effector Cell Lines | Stable, inducible expression of CRISPRa/i machinery. | Tet-On 3G dCas9-VPR or dCas9-KRAB lentiviral particles (Addgene #125755, #125756). |
| Paired sgRNA Libraries | Matched genome-wide libraries for activation and interference. | Calabrese CRISPRa and CRISPRi Human Whole Genome Libraries (Addgene #125763, #125764). |
| Lentiviral Packaging Mix | For production of sgRNA library virus. | Lenti-X Packaging Single Shots (Takara Bio #631275). |
| Selection Antibiotics | For stable cell line and sgRNA pool selection. | Puromycin dihydrochloride, Blasticidin S HCl. |
| NGS Library Prep Kit | For amplification and barcoding of sgRNA sequences from genomic DNA. | NEBNext Ultra II Q5 Master Mix (NEB #M0544). |
| Analysis Software | Robust, open-source pipeline for screen analysis. | MAGeCK (source on GitHub). |
| Validated Control sgRNAs | Non-targeting and targeting controls for normalization. | Non-Targeting Control sgRNA Pool (Synthego). |
Application Notes
CRISPR activation (CRISPRa) and interference (CRISPRi) screens have revolutionized functional genomics by enabling systematic, large-scale interrogation of gene function through targeted transcriptional perturbation. The following case studies highlight successful applications across three critical therapeutic areas, framed within the thesis that pooled CRISPRa/i screens are indispensable for mapping disease-relevant gene regulatory networks and identifying novel therapeutic targets.
Oncology: Identifying Synthetic Lethal Interactions in KRAS-Mutant Cancers
Objective: To discover genes whose transcriptional activation confers vulnerability in KRAS-driven non-small cell lung cancer (NSCLC), a cancer subtype with limited therapeutic options.
Screen Design: A genome-wide CRISPRa screen using the SunTag system was performed in a KRASG12C mutant NSCLC cell line (NCI-H358). The screen utilized the SAM (Synergistic Activation Mediator) library targeting ~23,000 genes. Cells were cultured under normal proliferation conditions, and sgRNA abundance was measured via NGS at baseline and after 14 population doublings.
Key Findings: Enriched sgRNAs pointed to genes whose activation inhibited proliferation. The top hit was the Tumor Protein P53 Inducible Protein 3 (TP53I3/AIG1), a known redox-sensitive enzyme. Activation of TP53I3 induced severe oxidative stress and ferroptosis selectively in KRAS-mutant cells. Quantitative data is summarized in Table 1.
Table 1: Top Hits from Oncology CRISPRa Screen in KRAS-Mutant NSCLC
| Gene Symbol | Log2 Fold Change (sgRNA Enrichment) | p-value | False Discovery Rate (FDR) | Proposed Mechanism |
|---|---|---|---|---|
| TP53I3 | +4.2 | 3.1e-8 | 7.2e-5 | Induces oxidative stress/ferroptosis |
| RFK | +3.8 | 2.4e-7 | 1.1e-4 | Depletes cellular NADPH pool |
| NOX4 | +3.5 | 5.7e-7 | 1.8e-4 | Generates reactive oxygen species (ROS) |
| SLC7A11 | -2.1* | 9.8e-6 | 0.002 | Repression sensitizes to ferroptosis |
*Negative log2FC indicates sgRNA depletion; gene repression is detrimental.
Neuroscience: Uncovering Modulators of Neuronal Resilience in Alzheimer's Disease Models
Objective: To identify genes that, when repressed, protect against beta-amyloid (Aβ) toxicity, a key pathological feature of Alzheimer's disease.
Screen Design: A focused CRISPRi screen (~5,000 genes) using dCas9-KRAB was conducted in human iPSC-derived glutamatergic neurons. Neurons were transduced with a lentiviral library and subsequently treated with oligomeric Aβ42. Cell viability was measured after 96 hours. sgRNA representation was sequenced to identify depleted sgRNAs (where repression enhanced survival) and enriched sgRNAs (where repression reduced survival).
Key Findings: Repression of the Serine/Threonine Kinase 11 (STK11/LKB1) gene significantly enhanced neuronal survival upon Aβ insult. Follow-up studies showed that STK11 repression attenuated Aβ-induced hyperexcitation and mitochondrial dysfunction by modulating AMPK signaling. Key data is in Table 2.
Table 2: Key Modulators from Neuroscience CRISPRi Screen Against Aβ Toxicity
| Gene Symbol | Log2 Fold Change | p-value | FDR | Effect of Repression |
|---|---|---|---|---|
| STK11 | -3.9 | 6.5e-9 | 3.3e-5 | Protective: Reduces neuronal hyperexcitation |
| PTK2B | -2.5 | 4.2e-6 | 0.001 | Protective: Lowers calcium influx |
| SORT1 | +2.8 | 1.8e-7 | 8.9e-5 | Detrimental: Reduces trophic support |
| PROX1 | -2.1 | 7.3e-6 | 0.002 | Protective: Role in glial modulation |
Immunology: Mapping Regulatory Networks in T-cell Exhaustion for Cancer Immunotherapy
Objective: To systematically discover transcriptional regulators that reverse T-cell exhaustion, a dysfunctional state of T-cells in chronic infection and cancer.
Screen Design: A custom CRISPRa screen targeting 1,500 transcription factors and epigenetic regulators was performed in primary human CD8+ T-cells. Exhaustion was induced by chronic antigen stimulation. The screen identified sgRNAs that, upon activation of their target gene, led to enhanced cytokine production (IL-2, IFNγ) and sustained proliferative capacity upon re-stimulation.
Key Findings: Activation of the BATF (Basic Leucine Zipper ATF-Like Transcription Factor) family member BATF3 potently rejuvenated exhausted T-cells. BATF3 overexpression reprogrammed the epigenetic landscape, suppressing exhaustion markers (e.g., PD-1, TIM-3) and promoting a memory-like phenotype. Quantitative outcomes are in Table 3.
Table 3: Top Transcriptional Regulators from Immunology CRISPRa Screen in T-cell Exhaustion
| Gene Symbol | Fold Increase in IL-2+ Cells | p-value (vs. Control) | Exhaustion Marker (PD-1) Mean Fluorescence (MFI) |
|---|---|---|---|
| BATF3 | 8.5x | 1.2e-10 | 1,205 (vs. 4,810 in control) |
| ID3 | 5.2x | 3.7e-8 | 2,150 |
| TOX2 | 0.4x* | 2.1e-6 | 5,890 |
| Control sgRNA | 1.0x | - | 4,810 |
Activation of *TOX2 further suppressed T-cell function.
Experimental Protocols
Protocol 1: Pooled Genome-wide CRISPRa Screen for Synthetic Lethality
Principle: Identify genes whose transcriptional activation selectively inhibits proliferation of a specific cancer genotype.
Materials: See "Research Reagent Solutions" below. Procedure:
- Library Lentivirus Production: HEK293T cells are co-transfected with the sgRNA SAM library plasmid, psPAX2, and pMD2.G using PEI. Virus-containing supernatant is collected at 48 and 72 hours, concentrated, and titered.
- Cell Line Transduction & Selection: Target cells (e.g., NCI-H358) are transduced at an MOI of ~0.3 to ensure single sgRNA integration. Puromycin selection (2 µg/mL) is applied 48 hours post-transduction for 7 days.
- Screen Execution: A minimum of 50 million transduced cells are maintained, representing at least 500x coverage of the library. Cells are passaged for ~14 population doublings. 20 million cells are harvested at Day 0 (baseline) and at the endpoint for genomic DNA extraction.
- sgRNA Amplification & Sequencing: sgRNA cassettes are PCR-amplified from genomic DNA (KAPA HiFi HotStart ReadyMix). A second PCR adds Illumina adapters and sample indexes. Products are pooled and sequenced on an Illumina NextSeq (75bp single-end).
- Data Analysis: Fastq files are aligned to the library reference using Bowtie2. sgRNA counts are normalized. Enrichment/depletion analysis is performed using MAGeCK or PinAPL-Py to calculate log2 fold changes and statistical significance.
Protocol 2: CRISPRi Screen in iPSC-Derived Neurons for Neuroprotection
Principle: Identify gene repressions that confer resistance to beta-amyloid oligomer toxicity.
Materials: iPSC-derived glutamatergic neurons, CRISPRi sgRNA library (lentiviral), synthetic Aβ42 oligomers, neuron-specific culture medium. Procedure:
- Neuronal Differentiation & Transduction: Human iPSCs are differentiated into pure glutamatergic neurons using a validated small-molecule protocol over 35 days. Neurons are transduced with the dCas9-KRAB-expressing lentivirus first, followed by the sgRNA library lentivirus at a low MOI (<0.5) in the presence of polybrene (8 µg/mL).
- Challenge & Selection: 7 days post-sgRNA transduction, neurons are treated with 5 µM of prepared oligomeric Aβ42 for 96 hours. Control wells receive vehicle.
- Cell Harvest & DNA Extraction: Surviving cells are gently lysed directly on the plate. Genomic DNA is extracted using a column-based method, ensuring high molecular weight.
- NGS Library Prep & Analysis: A two-step PCR amplifies the sgRNA region. Sequencing and analytical steps follow Protocol 1, with a focus on identifying significantly depleted sgRNAs in the Aβ-treated condition versus vehicle control.
Protocol 3: CRISPRa Screen in Primary Human Exhausted T-Cells
Principle: Discover gene activations that reverse the exhausted phenotype and restore effector function.
Materials: Primary human CD8+ T-cells from healthy donors, anti-CD3/CD28 activation beads, IL-2, custom CRISPRa sgRNA library (lentiviral), flow cytometry antibodies (anti-CD8, PD-1, TIM-3, IFNγ, IL-2). Procedure:
- T-cell Activation & Exhaustion: Isolated CD8+ T-cells are activated with Dynabeads (1:1 bead-to-cell ratio) and IL-2 (50 U/mL). Exhaustion is induced by maintaining stimulation and IL-2 for 10-14 days, with bead refreshing every 3-4 days.
- Transduction: Exhausted T-cells are transduced with the CRISPRa (SAM) sgRNA library lentivirus by spinfection (1000g, 90 mins, 32°C) in the presence of polybrene (8 µg/mL) and IL-2 (100 U/mL).
- Functional Sorting: 7 days post-transduction, cells are re-stimulated with PMA/Ionomycin in the presence of brefeldin A for 5 hours. Cells are stained for surface markers, fixed, permeabilized, and stained intracellularly for IFNγ and IL-2. The top ~10% of double-positive (IFNγ+IL-2+) cells and the bottom ~30% of cytokine-negative cells are isolated by FACS.
- Genomic DNA & Analysis: Genomic DNA is extracted separately from each sorted population. sgRNAs are amplified, sequenced, and analyzed to identify sgRNAs enriched in the functional (double-positive) population.
Pathway & Workflow Visualizations
Title: CRISPRa Screen Workflow for Synthetic Lethality in Cancer
Title: STK11 Repression Protects Neurons from Aβ Toxicity
Title: BATF3 Activation Reverses T-cell Exhaustion
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in CRISPRa/i Screens | Example/Supplier |
|---|---|---|
| dCas9-VP64/SunTag (CRISPRa) | Engineered dCas9 fused to transcriptional activation domains. Recruits multiple activators to the target promoter. | Addgene: #61425 (SAM/dCas9-VP64), #1000000074 (SunTag-VP64) |
| dCas9-KRAB (CRISPRi) | Engineered dCas9 fused to the Krüppel-associated box (KRAB) repression domain. Recruits repressive complexes to induce targeted gene silencing. | Addgene: #71237 (dCas9-KRAB) |
| Pooled sgRNA Library | Lentiviral plasmid library containing thousands of sgRNAs targeting genes of interest, each with a unique barcode. | Custom design (Twist Bioscience) or pre-built (e.g., Calabrese, Horlbeck libraries) |
| Lentiviral Packaging Plasmids | Plasmids (psPAX2, pMD2.G) providing viral structural and envelope proteins necessary to produce infectious lentiviral particles. | Addgene: #12260, #12259 |
| Next-Generation Sequencing Kit | For high-throughput sequencing of sgRNA barcodes from amplified genomic DNA to determine their abundance. | Illumina NextSeq 500/550 High Output Kit v2.5 (75 Cycles) |
| Bioinformatics Software (MAGeCK) | Computational tool specifically designed for analyzing CRISPR screen data. Calculates sgRNA enrichment/depletion and gene-level significance. | https://sourceforge.net/p/mageck/wiki/Home/ |
| Polybrene (Hexadimethrine Bromide) | A cationic polymer that enhances viral transduction efficiency by neutralizing charge repulsion between virions and the cell membrane. | Sigma-Aldrich, TR-1003-G |
| Puromycin Dihydrochloride | Aminonucleoside antibiotic used for the selection of cells successfully transduced with lentiviral vectors containing a puromycin resistance gene. | Thermo Fisher, A1113803 |
Best Practices for Reproducibility and Data Sharing in Functional Genomics
Functional genomics, particularly high-throughput CRISPR activation and inhibition (CRISPRa/i) screens, is a cornerstone of modern target discovery and validation in drug development. The complexity and scale of these experiments necessitate rigorous standards for reproducibility and transparent data sharing to ensure scientific integrity and accelerate translational research.
Foundational Principles & Pre-Experimental Planning
2.1. The FAIR and ALCOA Principles All data generated must adhere to the FAIR (Findable, Accessible, Interoperable, Reusable) and ALCOA (Attributable, Legible, Contemporaneous, Original, Accurate) principles. This is non-negotiable for regulatory-grade research.
2.2. Detailed Experimental Design Documentation Prior to initiating a screen, a comprehensive experimental plan must be documented and version-controlled. This includes:
- Biological Question & Hypothesis: Clearly stated.
- Cell Line Authentication: Method (e.g., STR profiling) and passage number used.
- CRISPR Library Details: Vendor, catalog number, version, and sequences.
- Screen Parameters: MOI, selection strategy (antibiotic, FACS), timepoints.
- Control Guides: Positive and negative controls, non-targeting guides.
- Replication Strategy: Number of biological and technical replicates.
- Primary Data Analysis Pipeline: Software and key parameters defined in advance.
Protocols & Application Notes
Protocol 1: Execution of a Pooled CRISPRa/i Screen with Reproducibility in Mind
- Aim: To perform a reproducible pooled CRISPRa or CRISPRi screen for gene essentiality or drug-gene interactions.
- Materials: See "Research Reagent Solutions" table.
- Detailed Workflow:
- Library Amplification & QC: Amplify the sgRNA plasmid library (e.g., Calabrese, Brunello) from glycerol stock. Perform deep sequencing on the amplified pool to confirm guide representation. Quantify precisely by fluorometry. Note: Aliquots for long-term storage.
- Lentiviral Production: Produce lentivirus in HEK293T cells using a 3rd generation packaging system. Titer virus using p24 ELISA or functional titration on target cells.
- Cell Transduction: Transduce target cells at a low MOI (~0.3) to ensure single-guide integration. Include a non-transduced control. Use puromycin selection (dose determined by kill curve) for 3-7 days.
- Screen Harvest & Sequencing: Harvest cells at the experimental endpoint (T-final). For the initial timepoint (T0), harvest at least 50 million cells or 500x library coverage 72 hours post-selection. Extract genomic DNA using a scalable method (e.g., Qiagen Blood & Cell Culture Maxi Kit). Perform a two-step PCR to amplify sgRNA inserts and attach Illumina sequencing adapters and sample barcodes. Use a minimum of 1000x guide coverage per sample for PCR input.
- Sequencing: Sequence on an Illumina platform (HiSeq/NovaSeq) to achieve a minimum read depth of 500 reads per guide.
Protocol 2: Quantitative Data Processing & Analysis Pipeline
- Aim: To standardize the analysis of raw sequencing data from CRISPR screens into gene-level scores.
- Input: Demultiplexed FASTQ files.
- Software: MAGeCK (v0.5.9+) or PinAPL-Py.
- Steps:
- Read Alignment & Counting: Use
mageck countto align reads to the library reference file and generate a raw count table. - Quality Control (QC): Generate QC plots: read distribution, Gini index, sample correlation. Discard samples with low correlation to replicates.
- Normalization & Beta-Score Calculation: Use
mageck testto normalize counts (median normalization) and calculate beta scores (gene essentiality) and p-values using a robust rank aggregation (RRA) or negative binomial model. For CRISPRi/a, positive beta scores indicate gene repression/activation effects. - Hit Calling: Define hits using a combined threshold (e.g., FDR < 0.05 & |beta| > 0.5). Compare results across all replicates.
- Read Alignment & Counting: Use
Data Sharing Standards & Deposition
All raw and processed data must be deposited in public repositories before publication.
Table 1: Mandatory Data Deposition Repositories
| Data Type | Recommended Repository | Required Metadata | Accession Format Example |
|---|---|---|---|
| Raw Sequencing Reads | Sequence Read Archive (SRA) | Library layout, platform, cell line | SRR1234567 |
| Processed Count Tables & Results | Gene Expression Omnibus (GEO) | Sample descriptions, protocol, analysis method | GSE123456 |
| CRISPR Screen Metadata | BioStudies or project-specific database | Full experimental design, library map, analysis code | S-BSST123 |
| Analysis Code & Scripts | GitHub, GitLab, or Zenodo | README with software versions, dependencies | DOI:10.5281/zenodo.1234567 |
Visualization of Workflows & Relationships
Diagram Title: End-to-End Reproducible CRISPR Screen Workflow
Diagram Title: FAIR Data Principles for Functional Genomics
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents & Materials for Reproducible CRISPRa/i Screens
| Item | Function & Critical Specification | Example Vendor/Catalog | Notes for Reproducibility |
|---|---|---|---|
| CRISPRa/i Library | Pre-designed sgRNA pool targeting genes of interest. | Addgene (e.g., Calabrese CRISPRa v1.1, Brunello CRISPRi) | Always record library lot number and version. Obtain full sequence map. |
| Lentiviral Packaging Mix | 3rd generation system for safe, high-titer virus production. | Dharmacon, Invitrogen | Use same system across experiments for consistent titers. |
| Validated Cell Line | Screening model with confirmed identity and genotype. | ATCC, DSMZ | Mandatory: STR profiling and mycoplasma testing report. Record passage number. |
| Selection Antibiotic | (e.g., Puromycin) for selecting transduced cells. | Thermo Fisher, Sigma | Critical: Perform kill curve for each new cell line batch. |
| NGS Library Prep Kit | For amplifying sgRNA inserts from genomic DNA. | Illumina, NEB | Use kits with high fidelity polymerase to minimize PCR bias. |
| Alignment & Analysis Software | Tool for quantifying guides and gene-level statistics. | MAGeCK, PinAPL-Py | Fix version numbers (e.g., use Conda/Docker) for exact reproducibility. |
| Data Repository Access | Platform for public data deposition. | NCBI SRA/GEO, ENA, Zenodo | Prepare metadata templates early in the project. |
Conclusion
CRISPRa and CRISPRi screens have matured into indispensable, complementary tools for the functional genomics toolkit, enabling precise, scalable interrogation of gene activation and repression phenotypes. By mastering the foundational principles, rigorous methodological execution, proactive troubleshooting, and robust validation outlined here, researchers can reliably uncover novel biological insights and therapeutic targets. The future of these technologies lies in enhanced specificity, in vivo screening applications, and integration with single-cell multi-omics. As these screens become more sophisticated, they will continue to accelerate the translation of genetic discoveries into clinical breakthroughs, fundamentally shaping the next era of biomedical research and drug development.